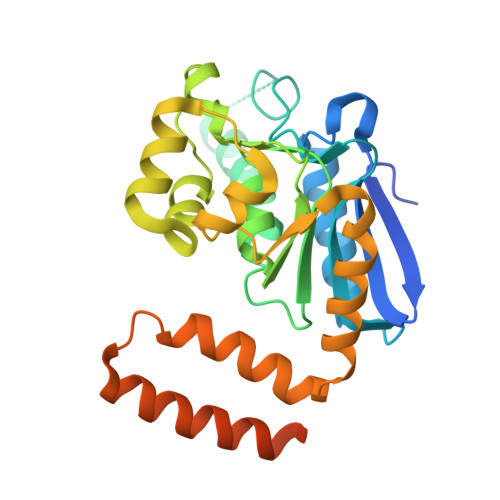
The structure of a prokaryotic feruloyl-CoA hydratase-lyase from a lignin-degrading consortium with high oligomerization stability under extreme pHs.
Liberato, M.V., Araujo, J.N., Sodre, V., Goncalves, T.A., Vilela, N., Moraes, E.C., Garcia, W., Squina, F.M.(2020) Biochim Biophys Acta Proteins Proteom 1868: 140344-140344
- PubMed: 31841665 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2019.140344
- Primary Citation Related Structures:
6P5U - PubMed Abstract:
In the context of increasing demand for renewable alternatives of fuels and chemicals, the valorization of lignin emerges as a value-adding strategy in biorefineries and an alternative to petroleum-derived molecules. One of the compounds derived from lignin is ferulic acid (FA), which can be converted into valuable molecules such as vanillin. In microorganisms, FA biotransformation into vanillin can occur via a two-step reaction catalyzed by the sequential activity of a feruloyl-CoA synthetase (FCS) and an feruloyl-CoA hydratase-lyase (FCHL), which could be exploited industrially. In this study, a prokaryotic FCHL derived from a lignin-degrading microbial consortium (named LM-FCHL) was cloned, successfully expressed in soluble form and purified. The crystal structure was solved and refined at 2.1 Å resolution. The LM-FCHL is a hexamer composed of a dimer of trimers, which showed to be quite stable under extreme pH conditions. Finally, small angle X-ray scattering corroborates the hexameric state in solution and indicates flexibility in the protein structure. The present study contributes to the field of lignin valorization to valuable molecules by establishing the biophysical and structural characterization for a novel FCHL member of unique characteristics.
- Programa de Processos Tecnológicos e Ambientais, Universidade de Sorocaba, Sorocaba, SP, Brazil.
Organizational Affiliation: