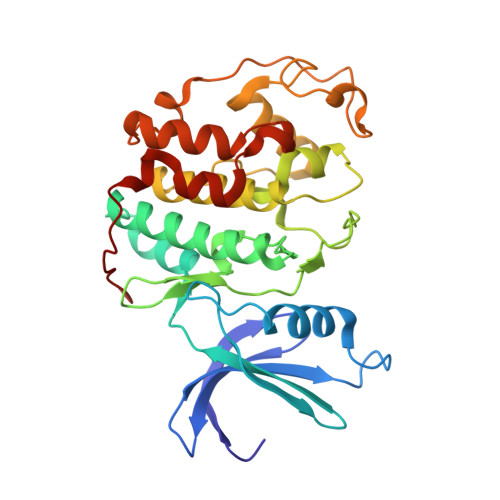
Structural basis for the ORC1-Cyclin A association.
Wang, B., Song, J.(2019) Protein Sci 28: 1727-1733
- PubMed: 31309634 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3689
- Primary Citation Related Structures:
6P3W - PubMed Abstract:
Progression of cell cycle is regulated by sequential expression of cyclins, which associate with distinct cyclin kinases to drive the transition between different cell cycle phases. The complex of Cyclin A with cyclin-dependent kinase 2 (CDK2) controls the DNA replication activity through phosphorylation of a set of chromatin factors, which critically influences the S phase transition. It has been shown that the direct interaction between the Cyclin A-CDK2 complex and origin recognition complex subunit 1 (ORC1) mediates the localization of ORC1 to centrosomes, where ORC1 inhibits cyclin E-mediated centrosome reduplication. However, the molecular basis underlying the specific recognition between ORC1 and cyclins remains elusive. Here we report the crystal structure of Cyclin A-CDK2 complex bound to a peptide derived from ORC1 at 2.54 å resolution. The structure revealed that the ORC1 peptide interacts with a hydrophobic groove, termed cyclin binding groove (CBG), of Cyclin A via a KXL motif. Distinct from other identified CBG-binding sequences, an arginine residue flanking the KXL motif of ORC1 inserts into a neighboring acidic pocket, contributing to the strong ORC1-Cyclin A association. Furthermore, structural and sequence analysis of cyclins reveals divergence on the ORC1-binding sites, which may underpin their differential ORC1-binding activities. This study provides a structural basis of the specific ORC1-cyclins recognition, with implication in development of novel inhibitors against the cyclin/CDK complexes.
- Department of Biochemistry, University of California, Riverside, California.
Organizational Affiliation: