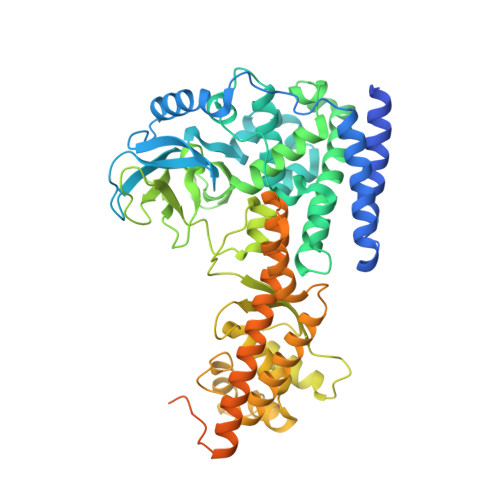
Structural basis for the target specificity of actin histidine methyltransferase SETD3.
Dai, S., Horton, J.R., Woodcock, C.B., Wilkinson, A.W., Zhang, X., Gozani, O., Cheng, X.(2019) Nat Commun 10: 3541-3541
- PubMed: 31388018 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-11554-6
- Primary Citation Related Structures:
6OX0, 6OX1, 6OX2, 6OX3, 6OX4, 6OX5 - PubMed Abstract:
SETD3 is an actin histidine-N 3 methyltransferase, whereas other characterized SET-domain enzymes are protein lysine methyltransferases. We report that in a pre-reactive complex SETD3 binds the N 3 -protonated form (N 3 -H) of actin His73, and in a post-reactive product complex, SETD3 generates the methylated histidine in an N 1 -protonated (N 1 -H) and N 3 -methylated form. During the reaction, the imidazole ring of His73 rotates ~105°, which shifts the proton from N 3 to N 1 , thus ensuring that the target atom N 3 is deprotonated prior to the methyl transfer. Under the conditions optimized for lysine deprotonation, SETD3 has weak lysine methylation activity on an actin peptide in which the target His73 is substituted by a lysine. The structure of SETD3 with Lys73-containing peptide reveals a bent conformation of Lys73, with its side chain aliphatic carbons tracing along the edge of imidazole ring and the terminal ε-amino group occupying a position nearly identical to the N 3 atom of unmethylated histidine.
- Department of Epigenetics and Molecular Carcinogenesis, The University of Texas MD Anderson Cancer Center, Houston, TX, 77030, USA.
Organizational Affiliation: