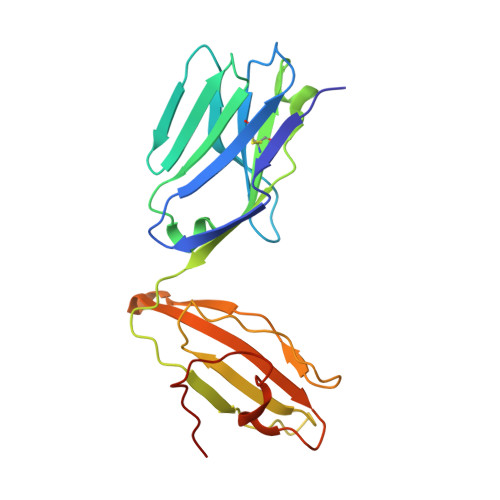
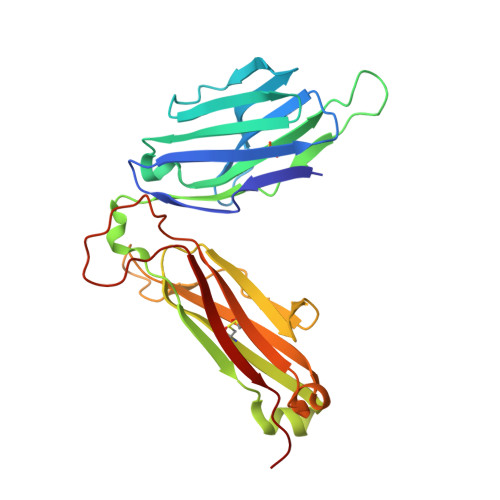
A TCR beta-Chain Motif Biases toward Recognition of Human CD1 Proteins.
Reinink, P., Shahine, A., Gras, S., Cheng, T.Y., Farquhar, R., Lopez, K., Suliman, S.A., Reijneveld, J.F., Le Nours, J., Tan, L.L., Leon, S.R., Jimenez, J., Calderon, R., Lecca, L., Murray, M.B., Rossjohn, J., Moody, D.B., Van Rhijn, I.(2019) J Immunol 203: 3395-3406
- PubMed: 31694911 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4049/jimmunol.1900872
- Primary Citation Related Structures:
6OVN, 6OVO - PubMed Abstract:
High-throughput TCR sequencing allows interrogation of the human TCR repertoire, potentially connecting TCR sequences to antigenic targets. Unlike the highly polymorphic MHC proteins, monomorphic Ag-presenting molecules such as MR1, CD1d, and CD1b present Ags to T cells with species-wide TCR motifs. CD1b tetramer studies and a survey of the 27 published CD1b-restricted TCRs demonstrated a TCR motif in humans defined by the TCR β-chain variable gene 4-1 (TRBV4-1) region. Unexpectedly, TRBV4-1 was involved in recognition of CD1b regardless of the chemical class of the carried lipid. Crystal structures of two CD1b-specific TRBV4-1 + TCRs show that germline-encoded residues in CDR1 and CDR3 regions of TRBV4-1-encoded sequences interact with each other and consolidate the surface of the TCR. Mutational studies identified a key positively charged residue in TRBV4-1 and a key negatively charged residue in CD1b that is shared with CD1c, which is also recognized by TRBV4-1 TCRs. These data show that one TCR V region can mediate a mechanism of recognition of two related monomorphic Ag-presenting molecules that does not rely on a defined lipid Ag.
- Department of Infectious Diseases and Immunology, Faculty of Veterinary Medicine, Utrecht University, 3584CL Utrecht, the Netherlands.
Organizational Affiliation: