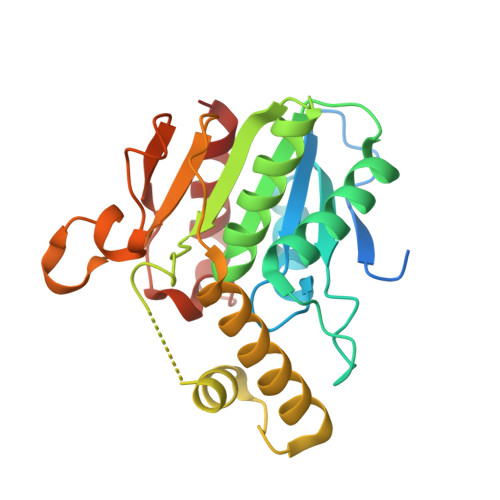
Structure of a bound peptide phosphonate reveals the mechanism of nocardicin bifunctional thioesterase epimerase-hydrolase half-reactions.
Patel, K.D., d'Andrea, F.B., Gaudelli, N.M., Buller, A.R., Townsend, C.A., Gulick, A.M.(2019) Nat Commun 10: 3868-3868
- PubMed: 31455765 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-11740-6
- Primary Citation Related Structures:
6OJC, 6OJD - PubMed Abstract:
Nonribosomal peptide synthetases (NRPSs) underlie the biosynthesis of many natural products that have important medicinal utility. Protection of the NRPS peptide products from proteolysis is critical to these pathways and is often achieved by structural modification, principally the introduction of D-amino acid residues into the elongating peptide. These amino acids are generally formed in situ from their L-stereoisomers by epimerization domains or dual-function condensation/epimerization domains. In singular contrast, the thioesterase domain of nocardicin biosynthesis mediates both the effectively complete L- to D-epimerization of its C-terminal amino acid residue (≥100:1) and hydrolytic product release. We report herein high-resolution crystal structures of the nocardicin thioesterase domain in ligand-free form and reacted with a structurally precise fluorophosphonate substrate mimic that identify the complete peptide binding pocket to accommodate both stereoisomers. These structures combined with additional functional studies provide detailed mechanistic insight into this unique dual-function NRPS domain.
- Department of Structural Biology, The Jacobs School of Medicine & Biomedical Sciences, State University of New York at Buffalo, 955 Main Street, Buffalo, NY, 14203, USA.
Organizational Affiliation: