Recognition Patterns of the C1/C2 Epitopes Involved in Fc-Mediated Response in HIV-1 Natural Infection and the RV114 Vaccine Trial.
Tolbert, W.D., Van, V., Sherburn, R., Tuyishime, M., Yan, F., Nguyen, D.N., Stanfield-Oakley, S., Easterhoff, D., Bonsignori, M., Haynes, B.F., Moody, M.A., Ray, K., Ferrari, G., Lewis, G.K., Pazgier, M.(2020) mBio 11
- PubMed: 32605979 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.00208-20
- Primary Citation Related Structures:
4FZ8, 6MG7, 6OED, 6OEJ, 6OFI - PubMed Abstract:
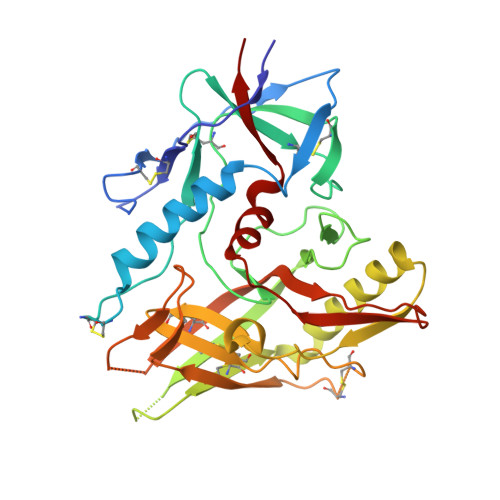
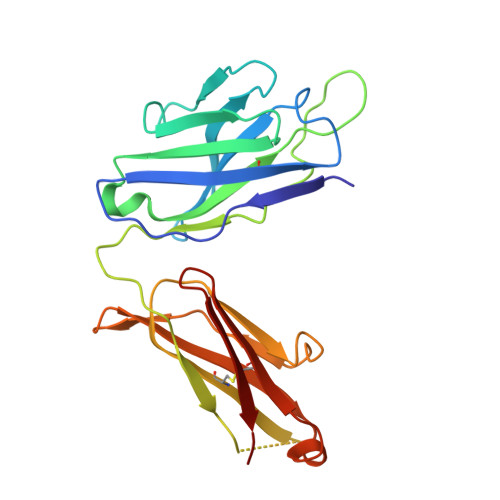
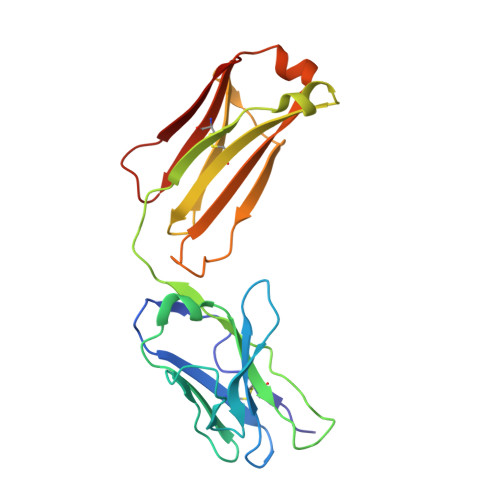
Antibodies (Abs) specific for CD4-induced envelope (Env) epitopes within constant region 1 and 2 (C1/C2) were induced in the RV144 vaccine trial, where antibody-dependent cellular cytotoxicity (ADCC) correlated with reduced risk of HIV-1 infection. We combined X-ray crystallography and fluorescence resonance energy transfer-fluorescence correlation spectroscopy to describe the molecular basis for epitopes of seven RV144 Abs and compared them to A32 and C11, C1/C2 Abs induced in HIV infection. Our data indicate that most vaccine Abs recognize the 7-stranded β-sandwich of gp120, a unique hybrid epitope bridging A32 and C11 binding sites. Although primarily directed at the 7-stranded β-sandwich, some accommodate the gp120 N terminus in C11-bound 8-stranded conformation and therefore recognize a broader range of CD4-triggered Env conformations. Our data also suggest that Abs of RV144 and RV305, the RV144 follow-up study, although likely initially induced by the ALVAC-HIV prime encoding full-length gp120, matured through boosting with truncated AIDSVAX gp120 variants. IMPORTANCE Antibody-dependent cellular cytotoxicity (ADCC) correlated with a reduced risk of infection from HIV-1 in the RV144 vaccine trial, the only HIV-1 vaccine trial to date to show any efficacy. Antibodies specific for CD4-induced envelope (Env) epitopes within constant region 1 and 2 (cluster A region) were induced in the RV144 trial and their ADCC activities were implicated in the vaccine efficacy. We present structural analyses of the antigen epitope targets of several RV144 antibodies specific for this region and C11, an antibody induced in natural infection, to show what the differences are in epitope specificities, mechanism of antigen recognition, and ADCC activities of antibodies induced by vaccination and during the course of HIV infection. Our data suggest that the truncated AIDSVAX gp120 variants used in the boost of the RV144 regimen may have shaped the vaccine response to this region, which could also have contributed to vaccine efficacy.
- Infectious Disease Division, Department of Medicine of Uniformed Services University of the Health Sciences, Bethesda, Maryland, USA.
Organizational Affiliation: