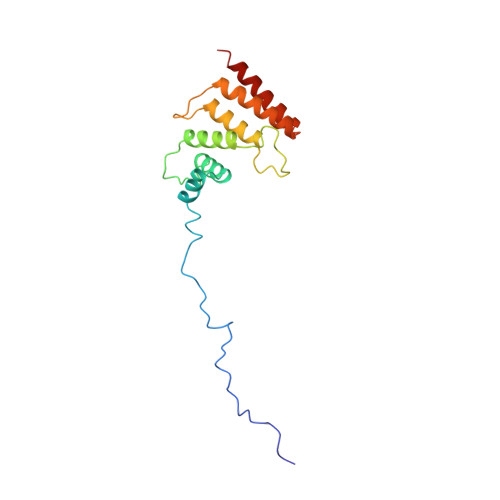
Solution Structure of the Carboxy-Terminal Tandem Repeat Domain of Eukaryotic Elongation Factor 2 Kinase and Its Role in Substrate Recognition.
Piserchio, A., Will, N., Giles, D.H., Hajredini, F., Dalby, K.N., Ghose, R.(2019) J Mol Biology 431: 2700-2717
- PubMed: 31108082 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2019.05.019
- Primary Citation Related Structures:
6NX4 - PubMed Abstract:
Eukaryotic elongation factor 2 kinase (eEF-2K), an atypical calmodulin-activated protein kinase, regulates translational elongation by phosphorylating its substrate, eukaryotic elongation factor 2 (eEF-2), thereby reducing its affinity for the ribosome. The activation and activity of eEF-2K are critical for survival under energy-deprived conditions and is implicated in a variety of essential physiological processes. Previous biochemical experiments have indicated that the binding site for the substrate eEF-2 is located in the C-terminal domain of eEF-2K, a region predicted to harbor several α-helical repeats. Here, using NMR methodology, we have determined the solution structure of a C-terminal fragment of eEF-2K, eEF-2K 562-725 that encodes two α-helical repeats. The structure of eEF-2K 562-725 shows signatures characteristic of TPR domains and of their SEL1-like sub-family. Furthermore, using the analyses of NMR spectral perturbations and ITC measurements, we have localized the eEF-2 binding site on eEF-2K 562-725 . We find that eEF-2K 562-725 engages eEF-2 with an affinity comparable to that of the full-length enzyme. Furthermore, eEF-2K 562-725 is able to inhibit the phosphorylation of eEF-2 by full-length eEF-2K in trans. Our present studies establish that eEF-2K 562-725 encodes the major elements necessary to enable the eEF-2K/eEF-2 interactions.
- Department of Chemistry and Biochemistry, The City College of New York, NewYork, NY 10031, USA.
Organizational Affiliation: