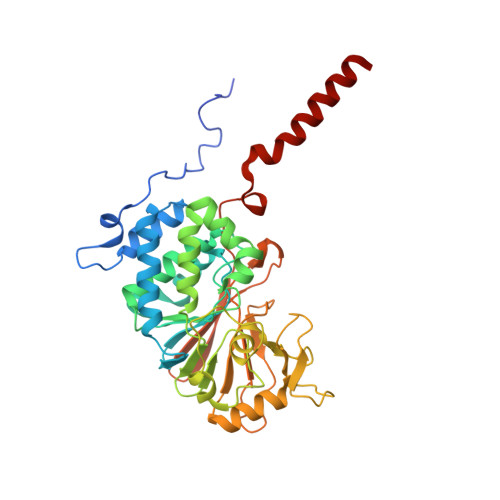
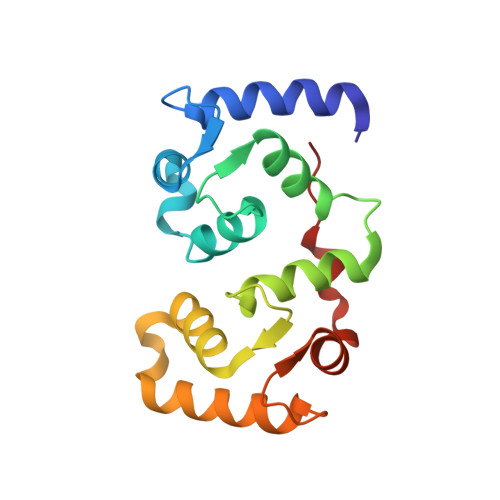
Molecular basis for the binding and selective dephosphorylation of Na+/H+exchanger 1 by calcineurin.
Hendus-Altenburger, R., Wang, X., Sjogaard-Frich, L.M., Pedraz-Cuesta, E., Sheftic, S.R., Bendsoe, A.H., Page, R., Kragelund, B.B., Pedersen, S.F., Peti, W.(2019) Nat Commun 10: 3489-3489
- PubMed: 31375679 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-11391-7
- Primary Citation Related Structures:
6NUC, 6NUF, 6NUU - PubMed Abstract:
Very little is known about how Ser/Thr protein phosphatases specifically recruit and dephosphorylate substrates. Here, we identify how the Na + /H + -exchanger 1 (NHE1), a key regulator of cellular pH homeostasis, is regulated by the Ser/Thr phosphatase calcineurin (CN). NHE1 activity is increased by phosphorylation of NHE1 residue T779, which is specifically dephosphorylated by CN. While it is known that Ser/Thr protein phosphatases prefer pThr over pSer, we show that this preference is not key to this exquisite CN selectivity. Rather a combination of molecular mechanisms, including recognition motifs, dynamic charge-charge interactions and a substrate interaction pocket lead to selective dephosphorylation of pT779. Our data identify T779 as a site regulating NHE1-mediated cellular acid extrusion and provides a molecular understanding of NHE1 substrate selection by CN, specifically, and how phosphatases recruit specific substrates, generally.
- Structural Biology and NMR Laboratory, Department of Biology, University of Copenhagen, Ole Maaløes Vej 5, DK-2200, Copenhagen N, Denmark.
Organizational Affiliation: