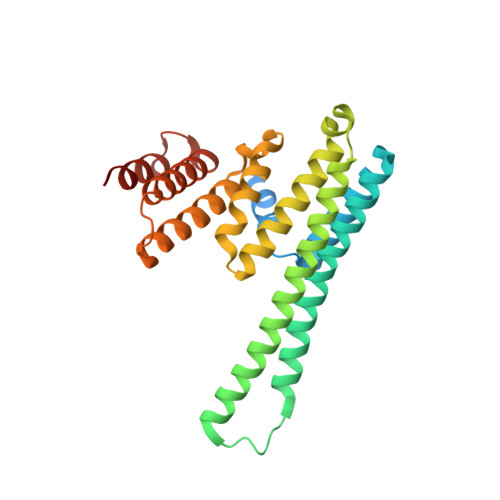
An asymmetric sheath controls flagellar supercoiling and motility in the Leptospira spirochete.
Gibson, K., Trajtenberg, F., Brady, M., San Martin, F., Mechaly, A., Wunder, E., Picardeau, M., Ko, A., Buschiazzo, A., Sindelar, C.(2020) Elife 9: e53672