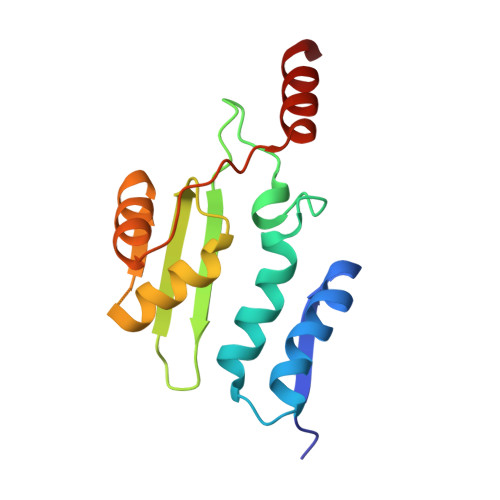
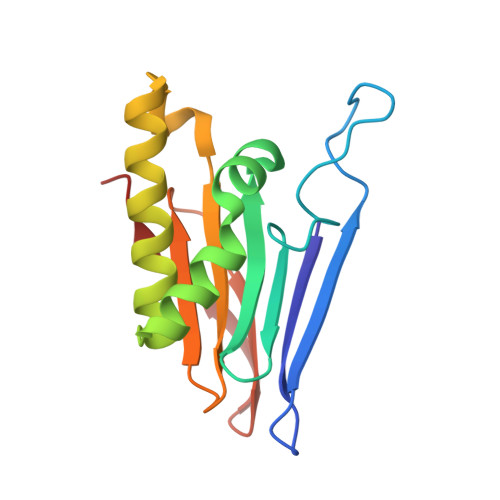
The T99K variant of glycosylasparaginase shows a new structural mechanism of the genetic disease aspartylglucosaminuria.
Pande, S., Guo, H.C.(2019) Protein Sci 28: 1013-1023
- PubMed: 30901125 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3607
- Primary Citation Related Structures:
6NQ6 - PubMed Abstract:
Aspartylglucosaminuria (AGU) is an inherited disease caused by mutations in a lysosomal amidase called aspartylglucosaminidase (AGA) or glycosylasparaginase (GA). This disorder results in an accumulation of glycoasparagines in the lysosomes of virtually all cell types, with severe clinical symptoms affecting the central nervous system, skeletal abnormalities, and connective tissue lesions. GA is synthesized as a single-chain precursor that requires an intramolecular autoprocessing to form a mature amidase. Previously, we showed that a Canadian AGU mutation disrupts this obligatory intramolecular autoprocessing with the enzyme trapped as an inactive precursor. Here, we report biochemical and structural characterization of a model enzyme corresponding to a new American AGU allele, the T99K variant. Unlike other variants with known 3D structures, this T99K model enzyme still has autoprocessing capacity to generate a mature form. However, its amidase activity to digest glycoasparagines remains low, consistent with its association with AGU. We have determined a 1.5-Å-resolution structure of this new AGU model enzyme and built an enzyme-substrate complex to provide a structural basis to analyze the negative effects of the T99K point mutation on K M and k cat of the amidase. It appears that a "molecular clamp" capable of fixing local disorders at the dimer interface might be able to rescue the deficiency of this new AGU variant.
- Department of Biological Sciences, University of Massachusetts Lowell, Lowell, Massachusetts, 01854.
Organizational Affiliation: