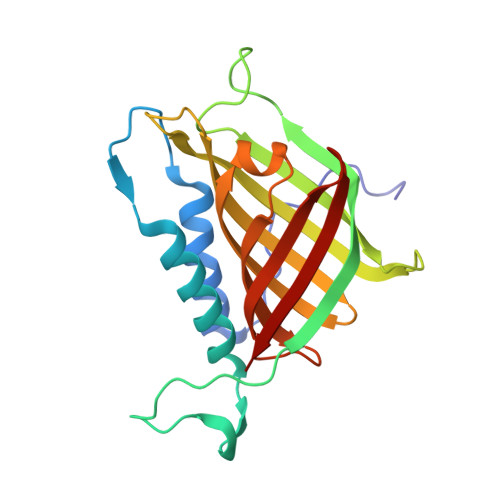
Unexpected enzyme-catalysed [4+2] cycloaddition and rearrangement in polyether antibiotic biosynthesis
Little, R., Paiva, F.C.R., Jenkins, R., Hong, H., Sun, Y., Demydchuk, Y., Samborskyy, M., Tosin, M., Leeper, F.J., Dias, M.V.B., Leadlay, P.F.(2019) Nat Catal