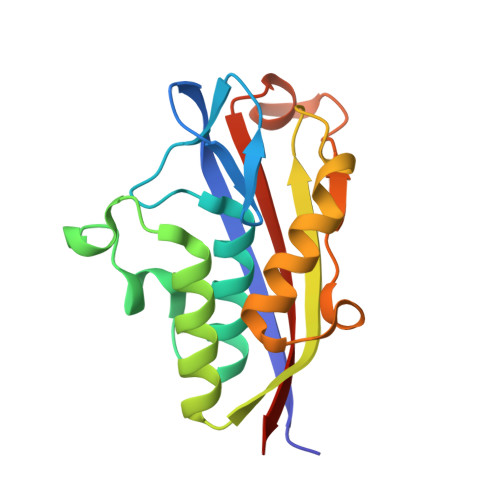
Crystal structure of 2C-methyl-D-erythritol 2,4-cyclodiphosphate synthase (IspF) Burkholderia pseudomallei in complex with ligand SR-4.
Abendroth, J., Dranow, D.M., Pierce, P.G., Rouffet, M., Lorimer, D.D., Horanyi, P.S., Edwards, T.E.To be published.