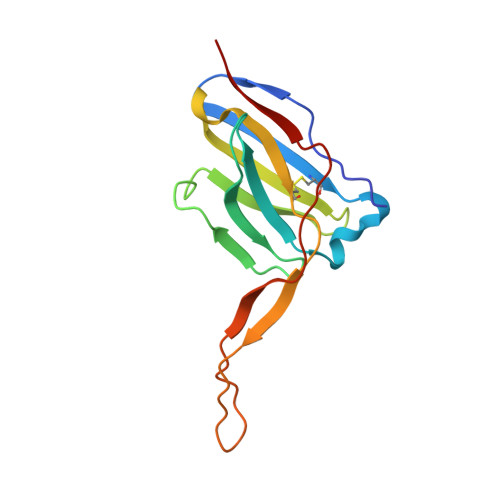
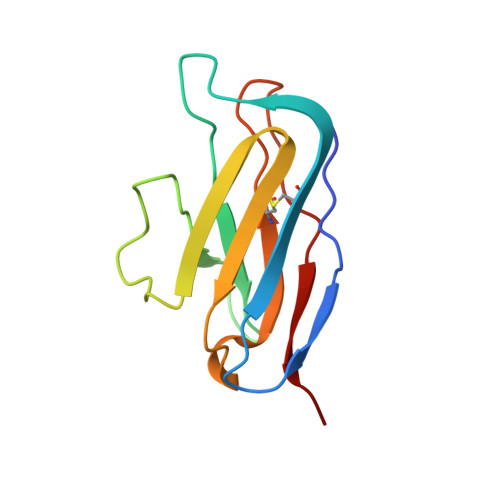
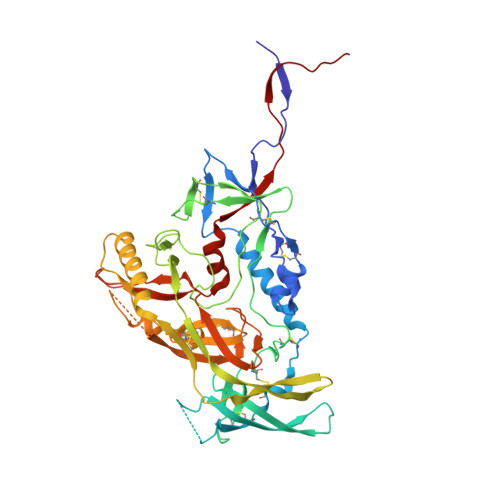
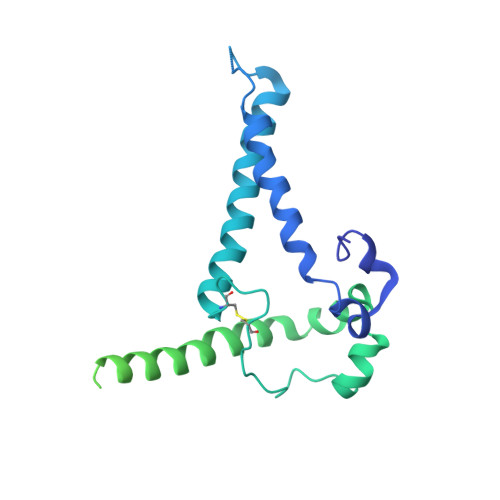
Similarities and differences between native HIV-1 envelope glycoprotein trimers and stabilized soluble trimer mimetics.
Torrents de la Pena, A., Rantalainen, K., Cottrell, C.A., Allen, J.D., van Gils, M.J., Torres, J.L., Crispin, M., Sanders, R.W., Ward, A.B.(2019) PLoS Pathog 15: e1007920-e1007920
- PubMed: 31306470 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1007920
- Primary Citation Related Structures:
6NIJ, 6OLP - PubMed Abstract:
The HIV-1 envelope glycoprotein (Env) trimer is located on the surface of the virus and is the target of broadly neutralizing antibodies (bNAbs). Recombinant native-like soluble Env trimer mimetics, such as SOSIP trimers, have taken a central role in HIV-1 vaccine research aimed at inducing bNAbs. We therefore performed a direct and thorough comparison of a full-length unmodified Env trimer containing the transmembrane domain and the cytoplasmic tail, with the sequence matched soluble SOSIP trimer, both based on an early Env sequence (AMC011) from an HIV+ individual that developed bNAbs. The structures of the full-length AMC011 trimer bound to either bNAb PGT145 or PGT151 were very similar to the structures of SOSIP trimers. Antigenically, the full-length and SOSIP trimers were comparable, but in contrast to the full-length trimer, the SOSIP trimer did not bind at all to non-neutralizing antibodies, most likely as a consequence of the intrinsic stabilization of the SOSIP trimer. Furthermore, the glycan composition of full-length and SOSIP trimers was similar overall, but the SOSIP trimer possessed slightly less complex and less extensively processed glycans, which may relate to the intrinsic stabilization as well as the absence of the membrane tether. These data provide insights into how to best use and improve membrane-associated full-length and soluble SOSIP HIV-1 Env trimers as immunogens.
- Department of Medical Microbiology, Amsterdam UMC - University of Amsterdam, Amsterdam, the Netherlands.
Organizational Affiliation: