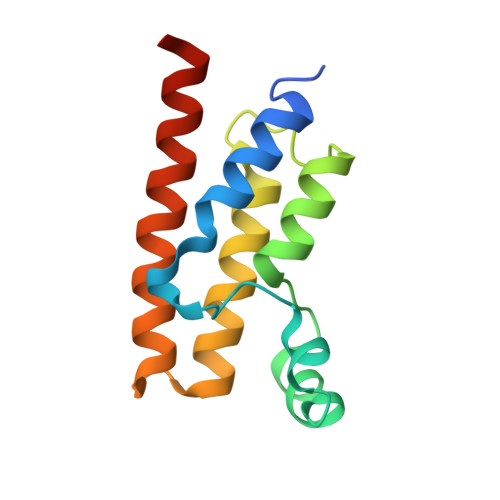
Trypanosoma brucei - BDF5, Tb427tmp.01.5000 A, solved with PF-CBP1
Lin, Y.H., Dong, A., Tempel, W., McAuley, J., Loppnau, P., Bountra, C., Arrowsmith, C.H., Edwards, A.M., Vedadi, M., Harding, R.J., Structural Genomics Consortium (SGC)To be published.