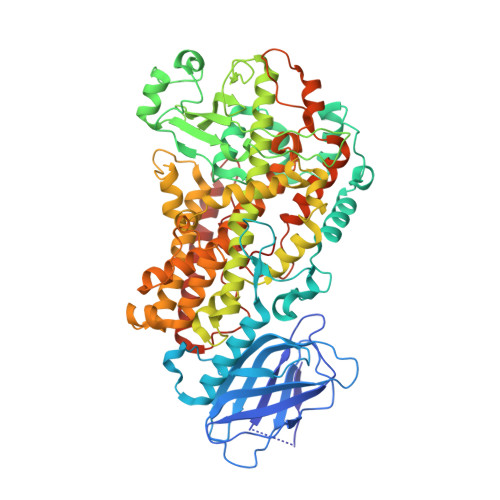
Structural and mechanistic insights into 5-lipoxygenase inhibition by natural products.
Gilbert, N.C., Gerstmeier, J., Schexnaydre, E.E., Borner, F., Garscha, U., Neau, D.B., Werz, O., Newcomer, M.E.(2020) Nat Chem Biol 16: 783-790
- PubMed: 32393899 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-020-0544-7
- Primary Citation Related Structures:
6N2W, 6NCF - PubMed Abstract:
Leukotrienes (LT) are lipid mediators of the inflammatory response that are linked to asthma and atherosclerosis. LT biosynthesis is initiated by 5-lipoxygenase (5-LOX) with the assistance of the substrate-binding 5-LOX-activating protein at the nuclear membrane. Here, we contrast the structural and functional consequences of the binding of two natural product inhibitors of 5-LOX. The redox-type inhibitor nordihydroguaiaretic acid (NDGA) is lodged in the 5-LOX active site, now fully exposed by disordering of the helix that caps it in the apo-enzyme. In contrast, the allosteric inhibitor 3-acetyl-11-keto-beta-boswellic acid (AKBA) from frankincense wedges between the membrane-binding and catalytic domains of 5-LOX, some 30 Å from the catalytic iron. While enzyme inhibition by NDGA is robust, AKBA promotes a shift in the regiospecificity, evident in human embryonic kidney 293 cells and in primary immune cells expressing 5-LOX. Our results suggest a new approach to isoform-specific 5-LOX inhibitor development through exploitation of an allosteric site in 5-LOX.
- Department of Biological Sciences, Louisiana State University, Baton Rouge, LA, USA.
Organizational Affiliation: