Sera Antibody Repertoire Analyses Reveal Mechanisms of Broad and Pandemic Strain Neutralizing Responses after Human Norovirus Vaccination.
Lindesmith, L.C., McDaniel, J.R., Changela, A., Verardi, R., Kerr, S.A., Costantini, V., Brewer-Jensen, P.D., Mallory, M.L., Voss, W.N., Boutz, D.R., Blazeck, J.J., Ippolito, G.C., Vinje, J., Kwong, P.D., Georgiou, G., Baric, R.S.(2019) Immunity 50: 1530-1541.e8
- PubMed: 31216462 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.immuni.2019.05.007
- Primary Citation Related Structures:
6N81, 6N8D - PubMed Abstract:
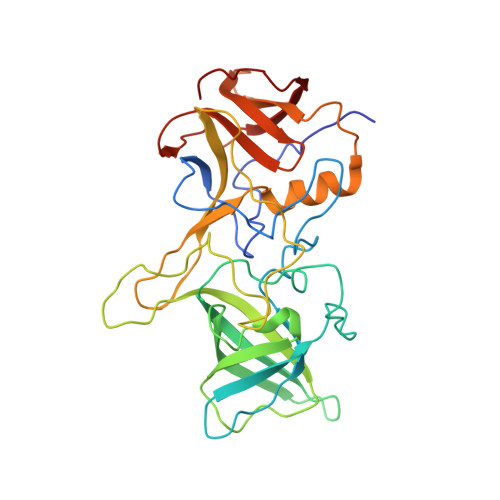
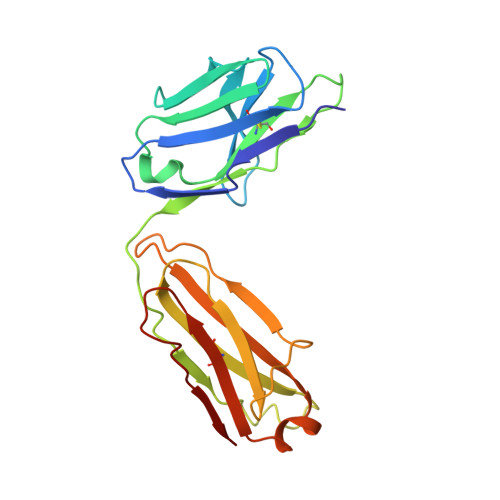
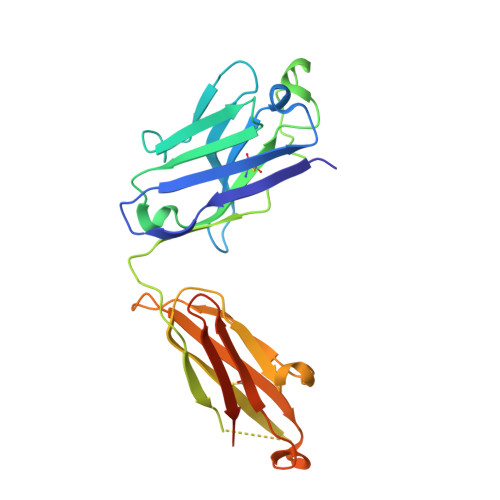
Rapidly evolving RNA viruses, such as the GII.4 strain of human norovirus (HuNoV), and their vaccines elicit complex serological responses associated with previous exposure. Specific correlates of protection, moreover, remain poorly understood. Here, we report the GII.4-serological antibody repertoire-pre- and post-vaccination-and select several antibody clonotypes for epitope and structural analysis. The humoral response was dominated by GII.4-specific antibodies that blocked ancestral strains or by antibodies that bound to divergent genotypes and did not block viral-entry-ligand interactions. However, one antibody, A1431, showed broad blockade toward tested GII.4 strains and neutralized the pandemic GII.P16-GII.4 Sydney strain. Structural mapping revealed conserved epitopes, which were occluded on the virion or partially exposed, allowing for broad blockade with neutralizing activity. Overall, our results provide high-resolution molecular information on humoral immune responses after HuNoV vaccination and demonstrate that infection-derived and vaccine-elicited antibodies can exhibit broad blockade and neutralization against this prevalent human pathogen.
- Department of Epidemiology, University of North Carolina, Chapel Hill, NC 27599, USA.
Organizational Affiliation: