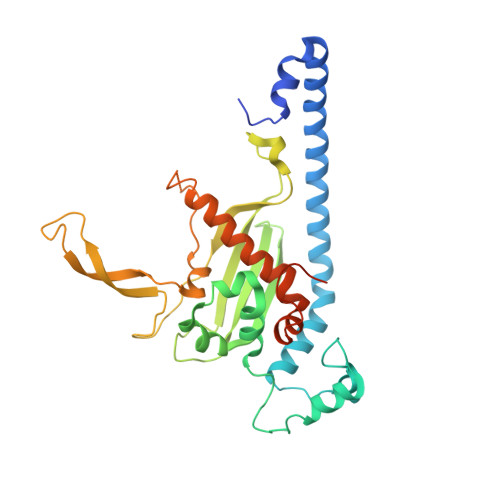
Characterization of Caenorhabditis elegans Nucleosome Assembly Protein 1 Uncovers the Role of Acidic Tails in Histone Binding.
Sarkar, P., Zhang, N., Bhattacharyya, S., Salvador, K., D'Arcy, S.(2019) Biochemistry 58: 108-113
- PubMed: 30521320 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.8b01033
- Primary Citation Related Structures:
6N2G - PubMed Abstract:
Nucleosome assembly proteins (Naps) influence chromatin dynamics by directly binding to histones. Here we provide a comprehensive structural and biochemical analysis of a Nap protein from Caenorhabditis elegans (CeNap1). CeNap1 naturally lacks the acidic N-terminal tail and has a short C-terminal tail compared to many other Nap proteins. Comparison of CeNap1 with full length and tail-less constructs of Saccharomyces cerevisiae Nap1 uncovers the role of these tails in self-association, histone binding, and Nap competition with DNA for H2A-H2B. We find that the presence of tails influences the stoichiometry of H2A-H2B binding and is required to complete the interactions between H2A-H2B and DNA. The absolute stoichiometry of the Nap protein and H2A-H2B complex is 2:1 or 2:2, with only a very small population of higher-order oligomers occurring at 150 mM NaCl. We also show that H3-H4 binds differently than H2A-H2B and that an (H3-H4) 2 tetramer can simultaneously bind two Nap 2 protein homodimers.
- Department of Biochemistry and Molecular Biology , The University of Melbourne , Melbourne 3000 , Australia.
Organizational Affiliation: