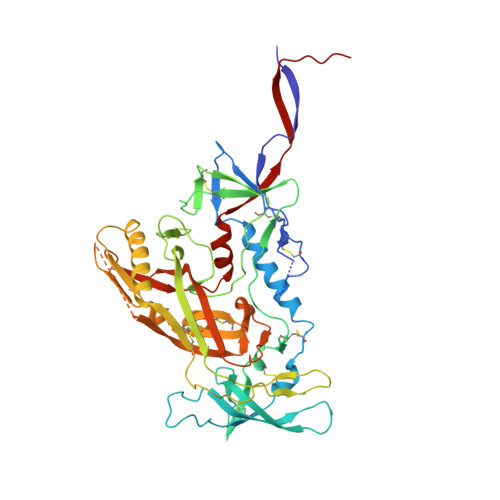
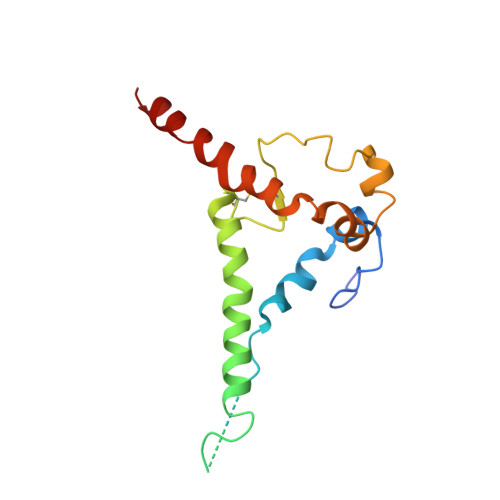
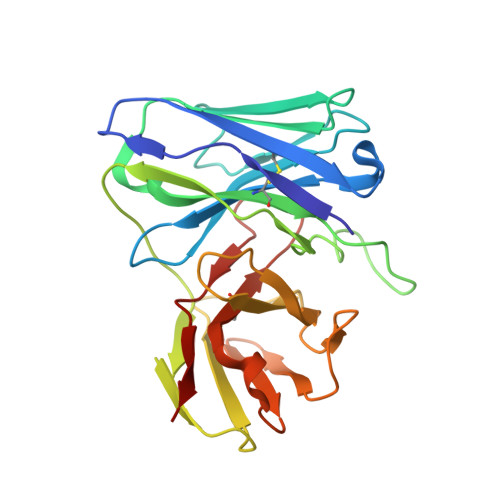
Kappa chain maturation helps drive rapid development of an infant HIV-1 broadly neutralizing antibody lineage.
Simonich, C.A., Doepker, L., Ralph, D., Williams, J.A., Dhar, A., Yaffe, Z., Gentles, L., Small, C.T., Oliver, B., Vigdorovich, V., Mangala Prasad, V., Nduati, R., Sather, D.N., Lee, K.K., Matsen Iv, F.A., Overbaugh, J.(2019) Nat Commun 10: 2190-2190
- PubMed: 31097697 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-09481-7
- Primary Citation Related Structures:
6MN7 - PubMed Abstract:
HIV-infected infants develop broadly neutralizing plasma responses with more rapid kinetics than adults, suggesting the ontogeny of infant responses could better inform a path to achievable vaccine targets. Here we reconstruct the developmental lineage of BF520.1, an infant-derived HIV-specific broadly neutralizing antibody (bnAb), using computational methods developed specifically for this purpose. We find that the BF520.1 inferred naive precursor binds HIV Env. We also show that heterologous cross-clade neutralizing activity evolved in the infant within six months of infection and that, ultimately, only 2% SHM is needed to achieve the full breadth of the mature antibody. Mutagenesis and structural analyses reveal that, for this infant bnAb, substitutions in the kappa chain were critical for activity, particularly in CDRL1. Overall, the developmental pathway of this infant antibody includes features distinct from adult antibodies, including several that may be amenable to better vaccine responses.
- Division of Human Biology, Fred Hutchinson Cancer Research Center, Seattle, WA, 98109, USA.
Organizational Affiliation: