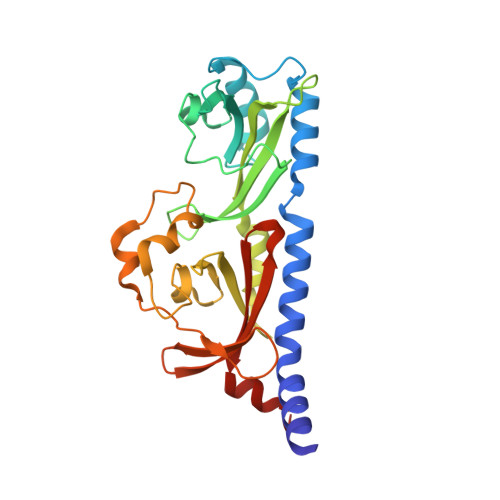
Structural and ligand binding analyses of the periplasmic sensor domain of RsbU in Chlamydia trachomatis support a role in TCA cycle regulation.
Soules, K.R., Dmitriev, A., LaBrie, S.D., Dimond, Z.E., May, B.H., Johnson, D.K., Zhang, Y., Battaile, K.P., Lovell, S., Hefty, P.S.(2020) Mol Microbiol 113: 68-88
- PubMed: 31637787 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/mmi.14401
- Primary Citation Related Structures:
6MAB - PubMed Abstract:
Chlamydia trachomatis is an obligate intracellular bacteria that undergo dynamic morphologic and physiologic conversions upon gaining an access to a eukaryotic cell. These conversions likely require the detection of key environmental conditions and regulation of metabolic activity. Chlamydia encodes homologs to proteins in the Rsb phosphoregulatory partner-switching pathway, best described in Bacillus subtilis. ORF CT588 has a strong sequence similarity to RsbU cytoplasmic phosphatase domain but also contains a unique periplasmic sensor domain that is expected to control the phosphatase activity. A 1.7 Å crystal structure of the periplasmic domain of the RsbU protein from C. trachomatis (PDB 6MAB) displays close structural similarity to DctB from Vibrio and Sinorhizobium. DctB has been shown, both structurally and functionally, to specifically bind to the tricarboxylic acid (TCA) cycle intermediate succinate. Surface plasmon resonance and differential scanning fluorimetry of TCA intermediates and potential metabolites from a virtual screen of RsbU revealed that alpha-ketoglutarate, malate and oxaloacetate bound to the RsbU periplasmic domain. Substitutions in the putative binding site resulted in reduced binding capabilities. An RsbU null mutant showed severe growth defects which could be restored through genetic complementation. Chemical inhibition of ATP synthesis by oxidative phosphorylation phenocopied the growth defect observed in the RsbU null strain. Altogether, these data support a model with the Rsb system responding differentially to TCA cycle intermediates to regulate metabolism and key differentiation processes.
- Department of Molecular Biosciences, University of Kansas, Lawrence, KS, 66045, USA.
Organizational Affiliation: