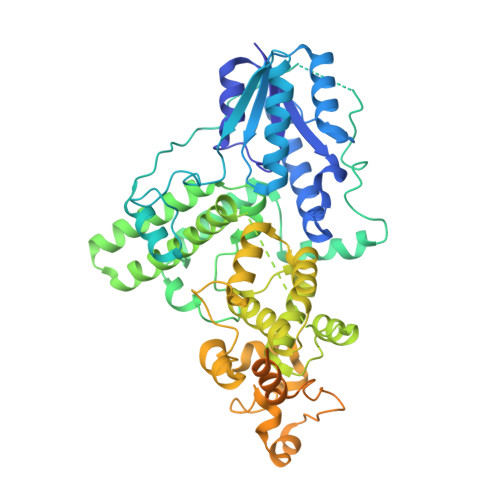
Structural insights into the photoactivation of Arabidopsis CRY2.
Ma, L., Guan, Z., Wang, Q., Yan, X., Wang, J., Wang, Z., Cao, J., Zhang, D., Gong, X., Yin, P.(2020) Nat Plants 6: 1432-1438
- PubMed: 33199893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41477-020-00800-1
- Primary Citation Related Structures:
6M79 - PubMed Abstract:
The blue-light receptor cryptochrome (CRY) in plants undergoes oligomerization to transduce blue-light signals after irradiation, but the corresponding molecular mechanism remains poorly understood. Here, we report the cryogenic electron microscopy structure of a blue-light-activated CRY2 tetramer at a resolution of 3.1 Å, which shows how the CRY2 tetramer assembles. Our study provides insights into blue-light-mediated activation of CRY2 and a theoretical basis for developing regulators of CRYs for optogenetic manipulation.
- National Key Laboratory of Crop Genetic Improvement and National Centre of Plant Gene Research, Huazhong Agricultural University, Wuhan, China.
Organizational Affiliation: