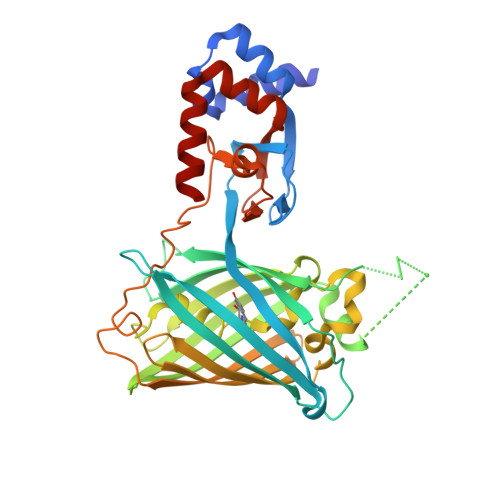
A high-performance genetically encoded fluorescent indicator for in vivo cAMP imaging.
Wang, L., Wu, C., Peng, W., Zhou, Z., Zeng, J., Li, X., Yang, Y., Yu, S., Zou, Y., Huang, M., Liu, C., Chen, Y., Li, Y., Ti, P., Liu, W., Gao, Y., Zheng, W., Zhong, H., Gao, S., Lu, Z., Ren, P.G., Ng, H.L., He, J., Chen, S., Xu, M., Li, Y., Chu, J.(2022) Nat Commun 13: 5363-5363
- PubMed: 36097007 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-32994-7
- Primary Citation Related Structures:
6M63 - PubMed Abstract:
cAMP is a key second messenger that regulates diverse cellular functions including neural plasticity. However, the spatiotemporal dynamics of intracellular cAMP in intact organisms are largely unknown due to low sensitivity and/or brightness of current genetically encoded fluorescent cAMP indicators. Here, we report the development of the new circularly permuted GFP (cpGFP)-based cAMP indicator G-Flamp1, which exhibits a large fluorescence increase (a maximum ΔF/F 0 of 1100% in HEK293T cells), decent brightness, appropriate affinity (a K d of 2.17 μM) and fast response kinetics (an association and dissociation half-time of 0.20 and 0.087 s, respectively). Furthermore, the crystal structure of the cAMP-bound G-Flamp1 reveals one linker connecting the cAMP-binding domain to cpGFP adopts a distorted β-strand conformation that may serve as a fluorescence modulation switch. We demonstrate that G-Flamp1 enables sensitive monitoring of endogenous cAMP signals in brain regions that are implicated in learning and motor control in living organisms such as fruit flies and mice.
- Research Center for Biomedical Optics and Molecular Imaging, Shenzhen Key Laboratory for Molecular Imaging, Guangdong Provincial Key Laboratory of Biomedical Optical Imaging Technology, Shenzhen Institute of Advanced Technology, Chinese Academy of Sciences, Shenzhen, 518055, China.
Organizational Affiliation: