The crystal structure of Ac-AChBP in complex with LvIA analogs reveals the mechanism of its selectivity towards different nAChR subtypes
Zhu, X.P., Pan, S., Wang, X.Q., Luo, S.L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Soluble acetylcholine receptor | A, C [auth B], E [auth D], G, I | 207 | Aplysia californica | Mutation(s): 0 | 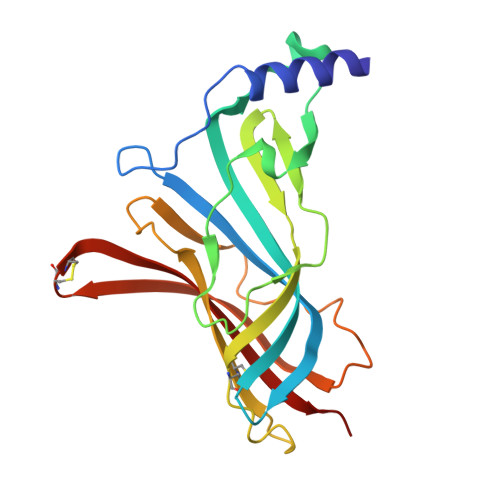 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8WSF8 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Alpha-conotoxin LvIA | B [auth F], D [auth C], F [auth E], H, J | 17 | Conus lividus | Mutation(s): 2 | 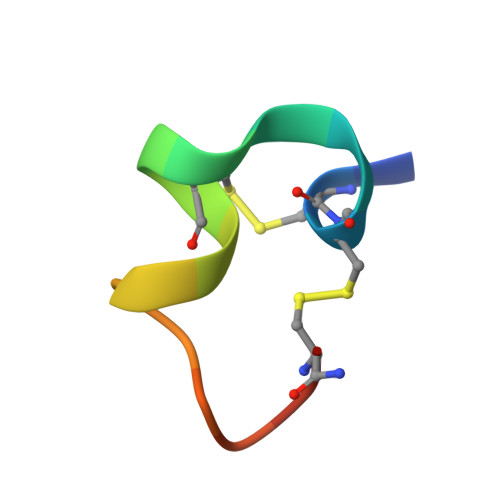 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | L8BU87 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 207.441 | α = 90 |
| b = 207.441 | β = 90 |
| c = 114.061 | γ = 90 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| PHASER | phasing |