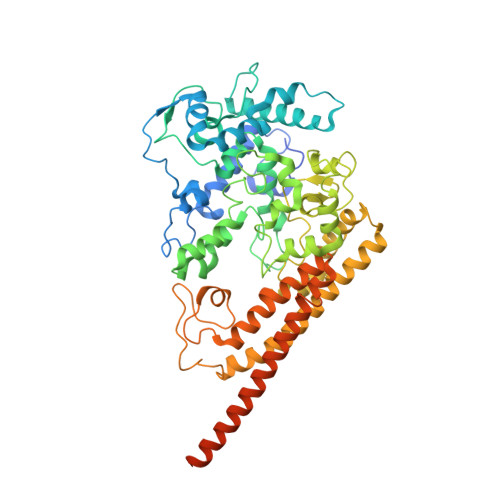
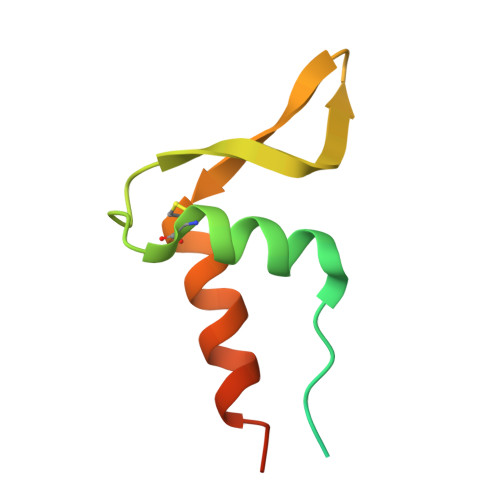
Juglone, a plant-derived 1,4-naphthoquinone, binds to hydroxylamine oxidoreductase and inhibits the electron transfer to cytochrome c 554.
Akutsu, Y., Fujiwara, T., Suzuki, R., Nishigaya, Y., Yamazaki, T.(2023) Appl Environ Microbiol 89: e0129123-e0129123
- PubMed: 38009977 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/aem.01291-23
- Primary Citation Related Structures:
6M0P, 6M0Q - PubMed Abstract:
Nitrification, the microbial conversion of ammonia to nitrate via nitrite, plays a pivotal role in the global nitrogen cycle. However, the excessive use of ammonium-based fertilizers in agriculture has disrupted this cycle, leading to groundwater pollution and greenhouse gas emissions. In this study, we have demonstrated the inhibitory effects of plant-derived juglone and related 1,4-naphthoquinones on the nitrification process in Nitrosomonas europaea . Notably, the inhibition mechanism is elucidated in which 1,4-naphthoquinones interact with hydroxylamine oxidoreductase, disrupting the electron transfer to cytochrome c 554 , a physiological electron acceptor. These findings support the notion that phytochemicals can impede nitrification by interfering with the essential electron transfer process in ammonia oxidation. The findings presented in this article offer valuable insights for the development of strategies aimed at the management of nitrification, reduction of fertilizer utilization, and mitigation of greenhouse gas emissions.
- Research Center for Advanced Analysis, National Agriculture and Food Research Organization (NARO), Tsukuba, Ibaraki, Japan.
Organizational Affiliation: