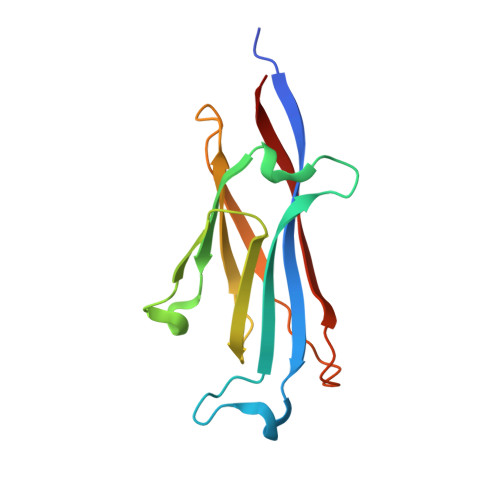
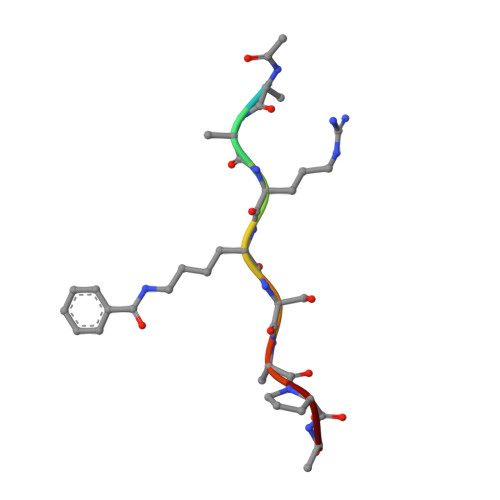
Histone benzoylation serves as an epigenetic mark for DPF and YEATS family proteins.
Ren, X., Zhou, Y., Xue, Z., Hao, N., Li, Y., Guo, X., Wang, D., Shi, X., Li, H.(2021) Nucleic Acids Res 49: 114-126
- PubMed: 33290558 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa1130
- Primary Citation Related Structures:
6LS6, 6LSB, 6LSD - PubMed Abstract:
Histone modifications and their functional readout serve as an important mechanism for gene regulation. Lysine benzoylation (Kbz) on histones is a recently identified acylation mark associated with active transcription. However, it remains to be explored whether putative readers exist to recognize this epigenetic mark. Here, our systematic binding studies demonstrated that the DPF and YEATS, but not the Bromodomain family members, are readers for histone Kbz. Co-crystal structural analyses revealed a 'hydrophobic encapsulation' and a 'tip-sensor' mechanism for Kbz readout by DPF and YEATS, respectively. Moreover, the DPF and YEATS family members display subtle yet unique features to create somewhat flexible engagements of different acylation marks. For instance, YEATS2 but not the other YEATS proteins exhibits best preference for Kbz than lysine acetylation and crotonylation due to its wider 'tip-sensor' pocket. The levels of histone benzoylation in cultured cells or in mice are upregulated upon sodium benzoate treatment, highlighting its dynamic regulation. In summary, our work identifies the first readers for histone Kbz and reveals the molecular basis underlying Kbz recognition, thus paving the way for further functional dissections of histone benzoylation.
- MOE Key Laboratory of Protein Sciences, Beijing Advanced Innovation Center for Structural Biology, Beijing Frontier Research Center for Biological Structure, Department of Basic Medical Sciences, School of Medicine, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: