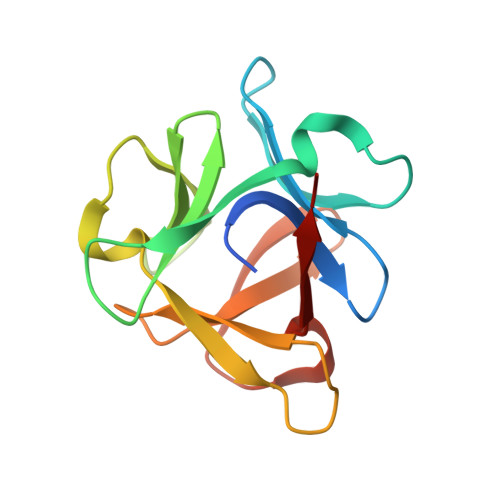
The structure of SeviL, a GM1b/asialo-GM1 binding R-type lectin from the mussel Mytilisepta virgata.
Kamata, K., Mizutani, K., Takahashi, K., Marchetti, R., Silipo, A., Addy, C., Park, S.Y., Fujii, Y., Fujita, H., Konuma, T., Ikegami, T., Ozeki, Y., Tame, J.R.H.(2020) Sci Rep 10: 22102-22102
- PubMed: 33328520 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-020-78926-7
- Primary Citation Related Structures:
6LF1, 6LF2 - PubMed Abstract:
SeviL is a recently isolated lectin found to bind to the linear saccharides of the ganglioside GM1b (Neu5Ac[Formula: see text](2-3)Gal[Formula: see text](1-3)GalNAc[Formula: see text](1-4)Gal[Formula: see text](1-4)Glc) and its precursor, asialo-GM1 (Gal[Formula: see text](1-3)GalNAc[Formula: see text](1-4)Gal[Formula: see text](1-4)Glc). The crystal structures of recombinant SeviL have been determined in the presence and absence of ligand. The protein belongs to the [Formula: see text]-trefoil family, but shows only weak sequence similarity to known structures. SeviL forms a dimer in solution, with one binding site per subunit, close to the subunit interface. Molecular details of glycan recognition by SeviL in solution were analysed by ligand- and protein-based NMR techniques as well as ligand binding assays. SeviL shows no interaction with GM1 due to steric hindrance with the sialic acid branch that is absent from GM1b. This unusual specificity makes SeviL of great interest for the detection and control of certain cancer cells, and cells of the immune system, that display asialo-GM1.
- Graduate School of Medical Life Science, Yokohama City University, 1-7-29 Suehiro, Yokohama, Kanagawa, 230-0045, Japan.
Organizational Affiliation: