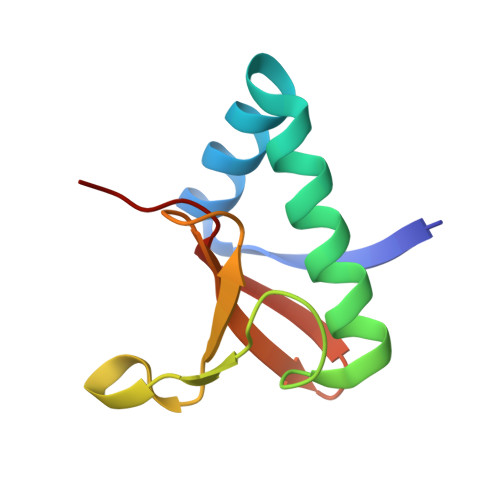
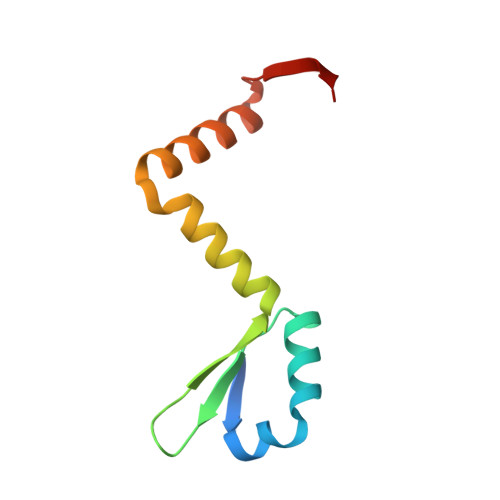
Distinct oligomeric structures of the YoeB-YefM complex provide insights into the conditional cooperativity of type II toxin-antitoxin system.
Xue, L., Yue, J., Ke, J., Khan, M.H., Wen, W., Sun, B., Zhu, Z., Niu, L.(2020) Nucleic Acids Res 48: 10527-10541
- PubMed: 32845304 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa706
- Primary Citation Related Structures:
6L8E, 6L8F, 7CUA - PubMed Abstract:
YoeB-YefM, the widespread type II toxin-antitoxin (TA) module, binds to its own promoter to autoregulate its transcription: repress or induce transcription under normal or stress conditions, respectively. It remains unclear how YoeB-YefM regulates its transcription depending on the YoeB to YefM TA ratio. We find that YoeB-YefM complex from S.aureus exists as two distinct oligomeric assemblies: heterotetramer (YoeB-YefM2-YoeB) and heterohexamer (YoeB-YefM2-YefM2-YoeB) with low and high DNA-binding affinities, respectively. Structures of the heterotetramer alone and heterohexamer bound to promoter DNA reveals that YefM C-terminal domain undergoes disorder to order transition upon YoeB binding, which allosterically affects the conformation of N-terminal DNA-binding domain. At TA ratio of 1:2, unsaturated binding of YoeB to the C-terminal regions of YefM dimer forms an optimal heterohexamer for DNA binding, and two YefM dimers with N-terminal domains dock into the adjacent major grooves of DNA to specifically recognize the 5'-TTGTACAN6AGTACAA-3' palindromic sequence, resulting in transcriptional repression. In contrast, at TA ratio of 1:1, binding of two additional YoeB molecules onto the heterohexamer induces the completely ordered conformation of YefM and disassembles the heterohexamer into two heterotetramers, which are unable to bind the promoter DNA optimally due to steric clashes, hence derepresses TA operon transcription.
- Hefei National Laboratory for Physical Sciences at the Microscale, Division of Molecular and Cellular Biophysics, University of Science and Technology of China, Hefei, Anhui 230026, China.
Organizational Affiliation: