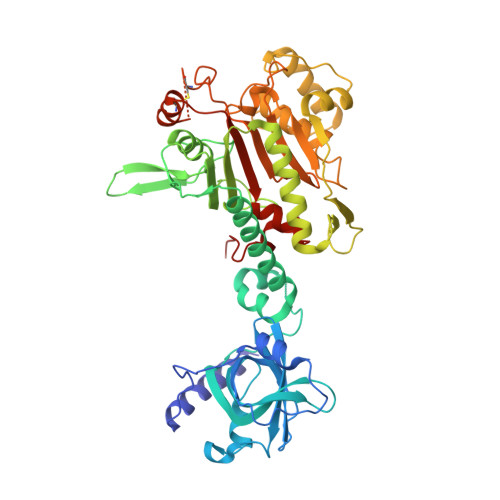
Crystal Structure of Lysyl-tRNA Synthetase from Plasmodium falciparum complexed with L-lysine and Cladosporin inhibitor, Cla-B
Babbar, P., Sato, M., Manickam, Y., Mishra, S., Harlos, K., Gupta, S., Parvez, S., Kikuchi, H., Sharma, A.(2021) Chembiochem