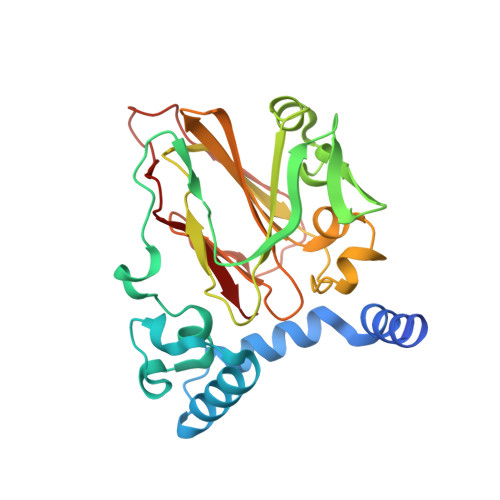
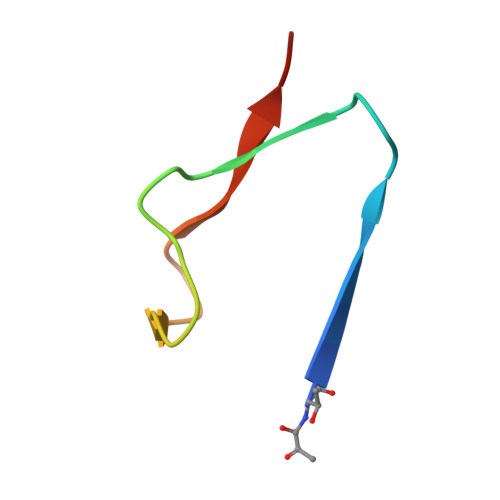
Structural Basis for Phosphatidylethanolamine Biosynthesis by Bacterial Phosphatidylserine Decarboxylase.
Watanabe, Y., Watanabe, Y., Watanabe, S.(2020) Structure 28: 799
- PubMed: 32402247 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2020.04.006
- Primary Citation Related Structures:
6L06, 6L07 - PubMed Abstract:
In both prokaryotes and eukaryotes, phosphatidylethanolamine (PE), one of the most abundant membrane phospholipids, plays important roles in various membrane functions and is synthesized through the decarboxylation of phosphatidylserine (PS) by PS decarboxylases (PSDs). However, the catalysis and substrate recognition mechanisms of PSDs remain unclear. In this study, we focused on the PSD from Escherichia coli (EcPsd) and determined the crystal structures of EcPsd in the apo form and PE-bound form at resolutions of 2.6 and 3.6 Å, respectively. EcPsd forms a homodimer, and each protomer has a positively charged substrate binding pocket at the active site. Structure-based mutational analyses revealed that conserved residues in the pocket are involved in PS decarboxylation. EcPsd has an N-terminal hydrophobic helical region that is important for membrane binding, thereby achieving efficient PS recognition. These results provide a structural basis for understanding the mechanism of PE biosynthesis by PSDs.
- Department of Bioscience, Graduate School of Agriculture, Ehime University, 3-5-7 Tarumi, Matsuyama, Ehime 790-8566, Japan. Electronic address: yasunori@agr.ehime-u.ac.jp.
Organizational Affiliation: