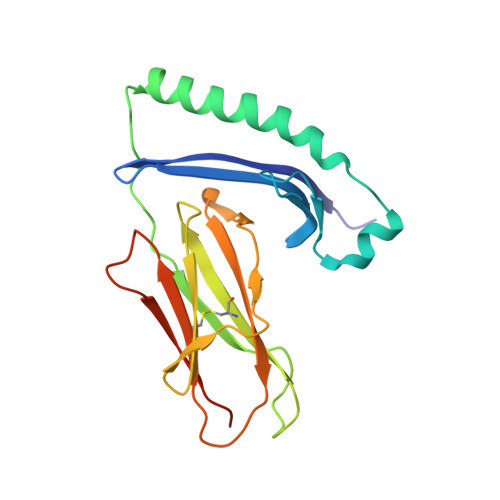
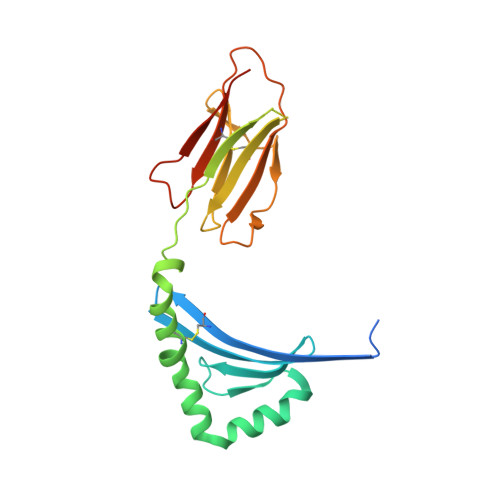
A Newly Recognized Pairing Mechanism of the alpha- and beta-Chains of the Chicken Peptide-MHC Class II Complex.
Zhang, L., Li, X., Ma, L., Zhang, B., Meng, G., Xia, C.(2020) J Immunol 204: 1630-1640
- PubMed: 32034060 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4049/jimmunol.1901305
- Primary Citation Related Structures:
6KVM - PubMed Abstract:
MHC class II (MHC-II) molecules play a crucial role in cellular and humoral immunity by forming peptide-MHC-II (pMHC-II) complexes. The three-dimensional structures of pMHC-II complexes have been well resolved in humans and mice. However, there is no structural information for pMHC-II complexes in nonmammals. In chickens, there are two closely related and highly polymorphic β-chains and one monomorphic α-chain, and the mechanism by which one monomorphic α-chain combines with two polymorphic β-chains to form a functional heterodimer remains unknown. In this study, we report the crystal structure of a chicken pMHC-II complex (pBL2*019:01) at 1.9-Å resolution as the first nonmammalian structure of a pMHC-II complex. The structure reveals an increase in hydrogen bonding between the α and β main chains at the central interface that is introduced by the insertion of four residues in the α-chain. The residues in the β-chain that form hydrogen bonds with the α-chain are conserved among all β alleles. These structural characteristics explain the phenomenon of only one BLA allele without sequence variation pairing with highly diverse BLB alleles from two loci in the genome. Additionally, the characteristics of the peptide in the peptide-binding groove were confirmed. These results provide a new understanding of the pairing mechanism of the α- and β-chains in a pMHC-II complex and establish a structural principle to design epitope-related vaccines for the prevention of chicken diseases.
- Department of Microbiology and Immunology, College of Veterinary Medicine, China Agricultural University, Haidian District, Beijing 100193, China; and.
Organizational Affiliation: