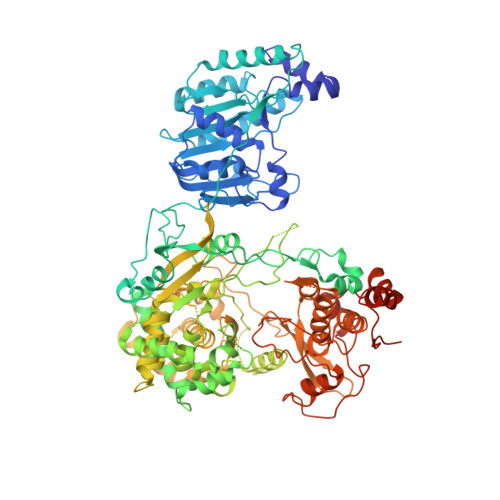
A conformation-based intra-molecular initiation factor identified in the flavivirus RNA-dependent RNA polymerase.
Wu, J., Ye, H.Q., Zhang, Q.Y., Lu, G., Zhang, B., Gong, P.(2020) PLoS Pathog 16: e1008484-e1008484
- PubMed: 32357182 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1008484
- Primary Citation Related Structures:
6KR2, 6KR3 - PubMed Abstract:
The flaviviruses pose serious threats to human health. Being a natural fusion of a methyltransferase (MTase) and an RNA-dependent RNA polymerase (RdRP), NS5 is the most conserved flavivirus protein and an important antiviral target. Previously reported NS5 structures represented by those from the Japanese encephalitis virus (JEV) and Dengue virus serotype 3 (DENV3) exhibit two apparently different global conformations, defining two sets of intra-molecular MTase-RdRP interactions. However, whether these NS5 conformations are conserved in flaviviruses and their specific functions remain elusive. Here we report two forms of DENV serotype 2 (DENV2) NS5 crystal structures representing two conformational states with defined analogies to the JEV-mode and DENV3-mode conformations, respectively, demonstrating the conservation of both conformation modes and providing clues for how different conformational states may be interconnected. Data from in vitro polymerase assays further demonstrate that perturbing the JEV-mode but not the DENV3-mode intra-molecular interactions inhibits catalysis only at initiation, while the cell-based virological analysis suggests that both modes of interactions are important for virus proliferation. Our work highlights the role of MTase as a unique intra-molecular initiation factor specifically only through the JEV-mode conformation, providing an example of conformation-based crosstalk between naturally fused protein functional modules.
- Key Laboratory of Special Pathogens and Biosafety, Wuhan Institute of Virology, Center for Biosafety Mega-Science, Chinese Academy of Sciences, Wuhan, Hubei, China.
Organizational Affiliation: