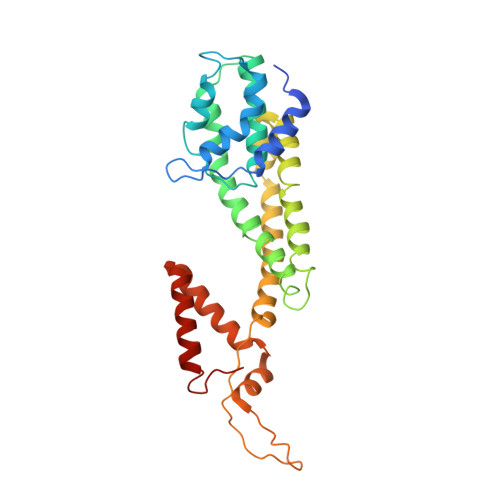
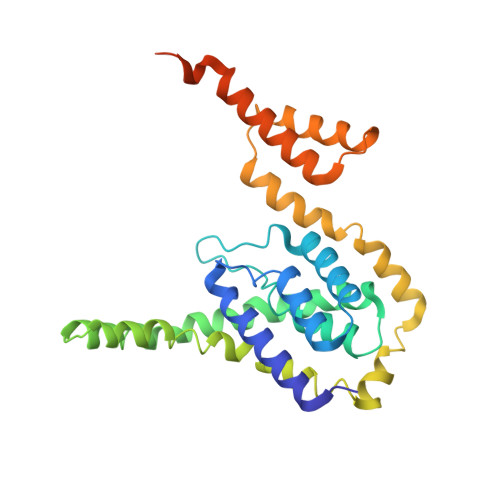
Structure of the mannose transporter of the bacterial phosphotransferase system.
Liu, X., Zeng, J., Huang, K., Wang, J.(2019) Cell Res 29: 680-682
- PubMed: 31209249 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-019-0194-z
- Primary Citation Related Structures:
6K1H - State Key Laboratory of Membrane Biology, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, 100084, Beijing, China.
Organizational Affiliation: