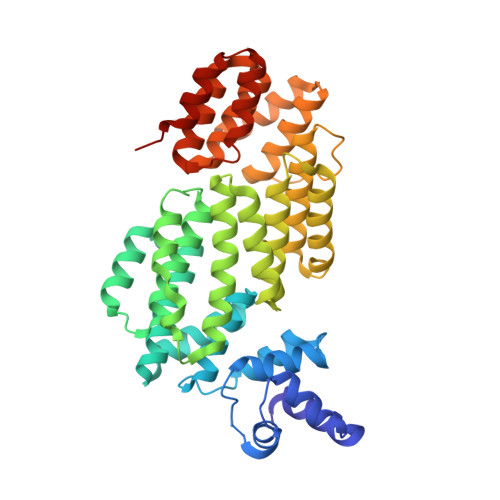
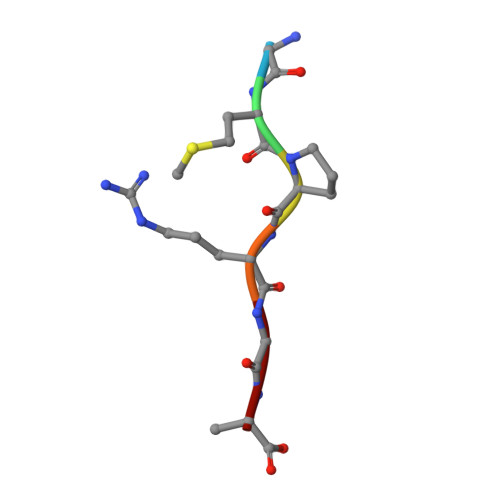
Structural insights into DNA recognition by AimR of the arbitrium communication system in the SPbeta phage.
Guan, Z.Y., Pei, K., Wang, J., Cui, Y.Q., Zhu, X., Su, X., Zhou, Y.B., Zhang, D.L., Tang, C., Yin, P., Liu, Z., Zou, T.T.(2019) Cell Discov 5: 29-29
- PubMed: 31149347 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41421-019-0101-2
- Primary Citation Related Structures:
6JG5, 6JG8, 6JG9 - PubMed Abstract:
A newly identified arbitrium communication system regulates the lysis-to-lysogeny decision in a Bacillus bacteriophage. This system contains an arbitrium hexapeptide as a signal, the cellular receptor AimR, and the lysogenic negative regulator AimX. AimR specifically targets the downstream DNA to activate aimX gene expression. The arbitrium peptide binds to AimR, inhibiting its DNA-binding to promote phage lysogeny. Recently, we and other groups have elucidated how arbitrium peptide sensed by AimR. However, the molecular mechanisms of DNA recognition by AimR and the regulation of its DNA-binding activity by the peptide remain largely unknown. Here, we report the crystal structure of the AimR-DNA complex at 2.1 Å resolution. The N-terminal HTH motif recognizes the palindromic DNA sequence, buttressed by interactions between positively charged residues and the DNA phosphate groups. The DNA-bound AimR assembles a more closed dimer than the peptide-bound form. Single-molecule FRET and crosslinking assays revealed that the AimR protein samples both open and closed conformations in solution. Arbitrium peptide binding induces a closed-to-open conformational change of AimR, eliminating DNA targeting. Our structural and functional analysis provides new insights into the DNA recognition mechanism of AimR and its regulation by the arbitrium peptide in the context of phage lysis-lysogeny decisions.
- 1National Key Laboratory of Crop Genetic Improvement and National Centre of Plant Gene Research, Huazhong Agricultural University, Wuhan, 430070 Hubei China.
Organizational Affiliation: