Design of an actin-severing peptide
Scipion, C.P.M., Robinson, R.C.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Actin, alpha skeletal muscle | 377 | Oryctolagus cuniculus | Mutation(s): 0 Gene Names: ACTA1, ACTA EC: 3.6.4 | 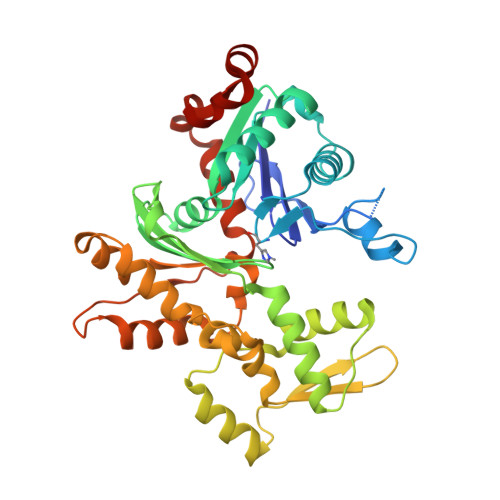 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P68135 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Peptide from Protein cordon-bleu | 22 | Mus musculus | Mutation(s): 1 Gene Names: Cobl, Kiaa0633 |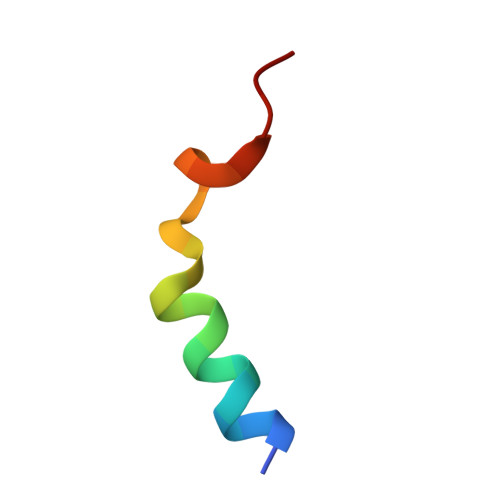  | |
UniProt & NIH Common Fund Data Resources | |||||
IMPC: MGI:105056 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q5NBX1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ATP Download:Ideal Coordinates CCD File | I [auth A], L [auth C], O [auth E], R [auth G] | ADENOSINE-5'-TRIPHOSPHATE C10 H16 N5 O13 P3 ZKHQWZAMYRWXGA-KQYNXXCUSA-N |  | ||
| CA Download:Ideal Coordinates CCD File | J [auth A] K [auth A] M [auth C] N [auth C] P [auth E] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| HIC Query on HIC | A, C, E, G | L-PEPTIDE LINKING | C7 H11 N3 O2 |  | HIS |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 58.22 | α = 90.62 |
| b = 81.192 | β = 107.19 |
| c = 89.869 | γ = 109.63 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHASER | phasing |