Crystal structure of Tyrosinase caddy protein(MelC1)with tyrosinase (MelC2)from Streptomyces avermitilis in complex with Zinc ion
Lee, S.-H., Hong, H., Kim, K.-J.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tyrosinase co-factor protein | 118 | Streptomyces avermitilis | Mutation(s): 0 Gene Names: melC1 | 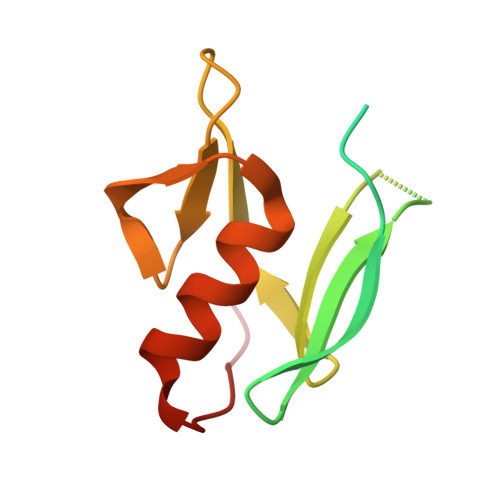 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q93HL1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tyrosinase | 285 | Streptomyces avermitilis | Mutation(s): 0 Gene Names: melC2 | 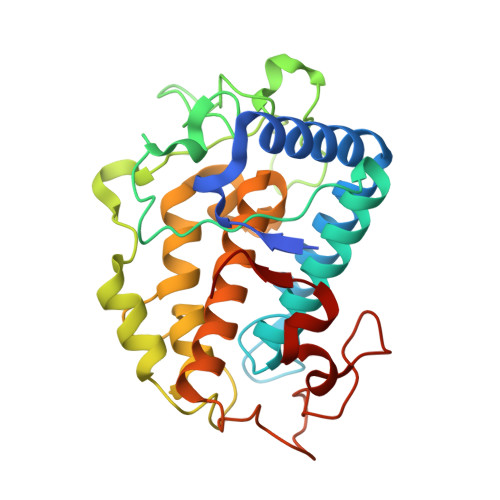 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q93HL2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ZN Download:Ideal Coordinates CCD File | C [auth B], D [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 119.998 | α = 90 |
| b = 71.828 | β = 113.72 |
| c = 48.524 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-2000 | data reduction |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data scaling |
| MOLREP | phasing |