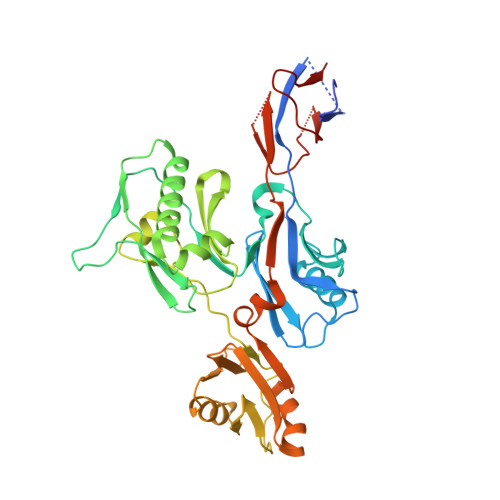
Structural analysis of the flagellar capping protein FliD from Helicobacter pylori.
Cho, S.Y., Song, W.S., Oh, H.B., Kim, H.U., Jung, H.S., Yoon, S.I.(2019) Biochem Biophys Res Commun 514: 98-104
- PubMed: 31023530 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.04.065
- Primary Citation Related Structures:
6IWY - PubMed Abstract:
Helicobacter pylori is a pathogenic flagellated bacterium that infects the gastroduodenal mucosa and causes peptic ulcers in humans. FliD caps the distal end of the flagellar filament and is essential in filament growth. Moreover, FliD has been studied to diagnose and prevent H. pylori infection. Here, we report structure-based molecular studies of H. pylori FliD (hpFliD). A crystal structure of hpFliD at 2.6 Å resolution presents a four-domain (D2-D5) structure, where the D3 domain forms a central platform surrounded by the other three domains (D2, D4, and D5). hpFliD domains D2 and D3 structurally resemble those of FliD orthologs, whereas the D4 and D5 domains are exclusive to hpFliD. Moreover, our ELISA analysis using anti-H. pylori antibodies demonstrated that the hpFliD-specific D4 and D5 domains are highly antigenic compared to the D2 and D3 domains. Collectively, our structural and serological analyses underscore the structural role of hpFliD domains and provide a molecular basis for vaccine and diagnosis development.
- Division of Biomedical Convergence, College of Biomedical Science, Kangwon National University, Chuncheon, 24341, Republic of Korea.
Organizational Affiliation: