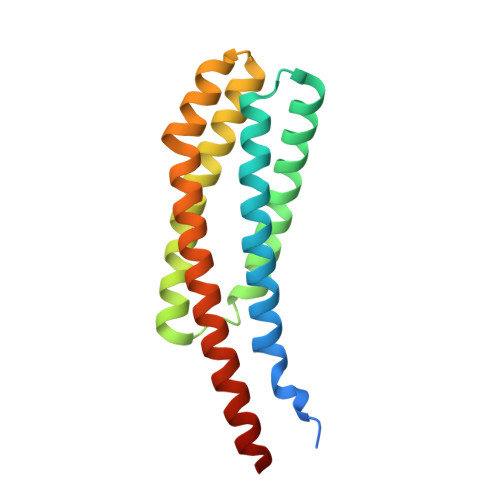
The ligand-binding domain of a chemoreceptor from Comamonas testosteroni has a previously unknown homotrimeric structure.
Hong, Y., Huang, Z., Guo, L., Ni, B., Jiang, C.Y., Li, X.J., Hou, Y.J., Yang, W.S., Wang, D.C., Zhulin, I.B., Liu, S.J., Li, D.F.(2019) Mol Microbiol 112: 906-917
- PubMed: 31177588 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/mmi.14326
- Primary Citation Related Structures:
5XUA, 5XUB, 6ITS - PubMed Abstract:
Transmembrane chemoreceptors are widely present in Bacteria and Archaea. They play a critical role in sensing various signals outside and transmitting to the cell interior. Here, we report the structure of the periplasmic ligand-binding domain (LBD) of the transmembrane chemoreceptor MCP2201, which governs chemotaxis to citrate and other organic compounds in Comamonas testosteroni. The apo-form LBD crystal revealed a typical four-helix bundle homodimer, similar to previously well-studied chemoreceptors such as Tar and Tsr of Escherichia coli. However, the citrate-bound LBD revealed a four-helix bundle homotrimer that had not been observed in bacterial chemoreceptor LBDs. This homotrimer was further confirmed with size-exclusion chromatography, analytical ultracentrifugation and cross-linking experiments. The physiological importance of the homotrimer for chemotaxis was demonstrated with site-directed mutations of key amino acid residues in C. testosteroni mutants.
- State Key Laboratory of Microbial Resources, Institute of Microbiology, Chinese Academy of Sciences, No. 1 Beichen West Road, Chaoyang District, Beijing, 100101, China.
Organizational Affiliation: