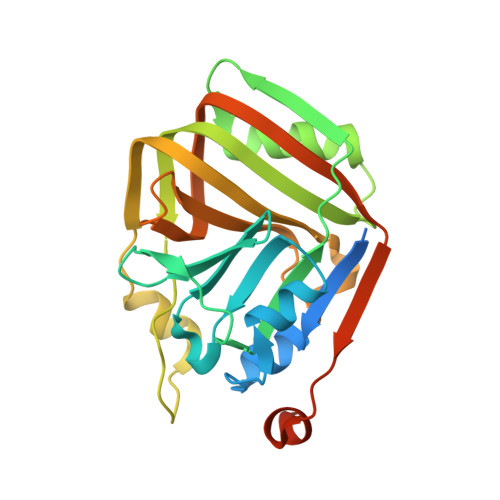
Structural characterization of an acetolactate decarboxylase from Klebsiella pneumoniae
Wu, W., Zhao, Q., Che, S., Jia, H., Liang, H., Zhang, H., Liu, R., Zhang, Q., Bartlam, M.(2019) Biochem Biophys Res Commun 509: 154-160
- PubMed: 30580999 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2018.12.094
- Primary Citation Related Structures:
6INB, 6INC - PubMed Abstract:
Acetolactate decarboxylase (ALDC) is a well-characterized anabolic enzyme involved with 3-hydroxy butanone (acetoin), an important physiological metabolite excreted by microbes. Although the enzyme is widely present in microorganisms, few atomic structures and functions of ALDC have been reported to date. Here we report the crystal structure of ALDC from Klebsiella pneumoniae KP (KpALDC). KpALDC crystallizes in space group P3 1 21 with one monomer per asymmetric unit. Analytical ultracentrifugation data shows that KpALDC forms a stable dimer but can exist as a tetramer in solution. A Zn 2+ ion is coordinated by three strictly-conserved histidines (His198, His200 and His211) and a conserved glutamate (Glu69), but the C-terminal tail that forms part of the active site in ALDC enzymes is disordered. A complex structure with ethane-1,2-diol shows a unusual mode of binding, whereby the ligand does not coordinate the Zn 2+ ion. MicroScale Thermophoresis analysis shows that KpALDC binds Zn 2+ ions, but no binding of Mg 2+ , Ca 2+ and Mn 2+ ions was detected.
- State Key Laboratory of Medicinal Chemical Biology, Nankai University, Tianjin, 300071, China; College of Life Sciences, Nankai University, Tianjin, 300071, China.
Organizational Affiliation: