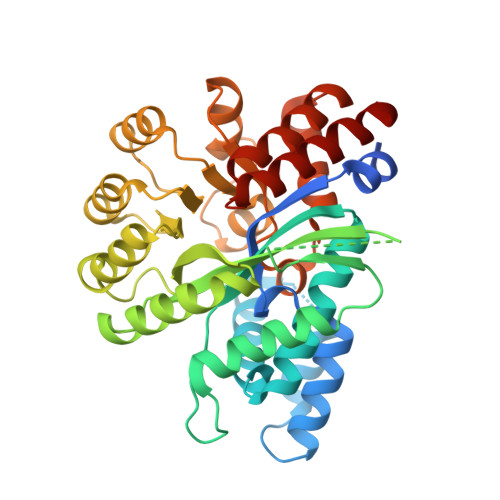
Alternative conformation induced by substrate binding for Arabidopsis thalianaN6-methyl-AMP deaminase.
Jia, Q., Xie, W.(2019) Nucleic Acids Res 47: 3233-3243
- PubMed: 30721978 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkz070
- Primary Citation Related Structures:
6IJM, 6IJN, 6IJP - PubMed Abstract:
Adenosine deaminase is involved in adenosine degradation and salvage pathway, and plays important physiological roles in purine metabolism. Recently, a novel type of adenosine deaminase-like protein has been identified, which displays deamination activity toward N6-methyl-adenosine monophosphate but not adenosine or AMP, and was consequently named N6-methyl-AMP deaminase (MAPDA). The underlying structural basis of MAPDA recognition and catalysis is poorly understood. Here, we present the crystal structures of MAPDA from Arabidopsis thaliana in the free and in the ligand-bound forms. The protein contains a conserved (β/α)8 Tim-barrel domain and a typical zinc-binding site, but it also exhibits idiosyncratic local differences for two flexible helices important for substrate binding. The extensive interactions between the N6-methyl-AMP substrate or the inosine monophosphate product and the enzyme were identified, and subsequently evaluated by the deamination activity assays. Importantly, each structure reported here represents a different stage of the catalytic pathway and their structural differences suggested that the enzyme can exist in two distinct conformational states. The open state switches to the closed one upon the binding of ligands, brought about by the two critical helices. Our structural studies provide the first look of this important metabolic enzyme and shed lights on its catalytic pathway, which holds promise for the structure-based drug design for MAPDA-related diseases.
- MOE Key Laboratory of Gene Function and Regulation, State Key Laboratory for Biocontrol, School of Life Sciences, The Sun Yat-Sen University, Guangzhou, Guangdong, 510006, People's Republic of China.
Organizational Affiliation: