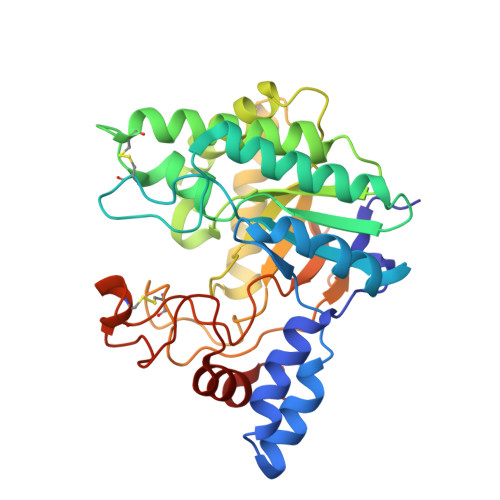
Crystal structures of the GH6 Orpinomyces sp. Y102 CelC7 enzyme with exo and endo activity and its complex with cellobiose.
Huang, H.C., Qi, L.H., Chen, Y.C., Tsai, L.C.(2019) Acta Crystallogr D Struct Biol 75: 1138-1147
- PubMed: 31793907 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798319013597
- Primary Citation Related Structures:
6IDW - PubMed Abstract:
The catalytic domain (residues 128-449) of the Orpinomyces sp. Y102 CelC7 enzyme (Orp CelC7) exhibits cellobiohydrolase and cellotriohydrolase activities. Crystal structures of Orp CelC7 and its cellobiose-bound complex have been solved at resolutions of 1.80 and 2.78 Å, respectively. Cellobiose occupies subsites +1 and +2 within the active site of Orp CelC7 and forms hydrogen bonds to two key residues: Asp248 and Asp409. Furthermore, its substrate-binding sites have both tunnel-like and open-cleft conformations, suggesting that the glycoside hydrolase family 6 (GH6) Orp CelC7 enzyme may perform enzymatic hydrolysis in the same way as endoglucanases and cellobiohydrolases. LC-MS/MS analysis revealed cellobiose (major) and cellotriose (minor) to be the respective products of endo and exo activity of the GH6 Orp CelC7.
- Institute of Organic and Polymeric Materials, National Taipei University of Technology, Taipei, Taiwan.
Organizational Affiliation: