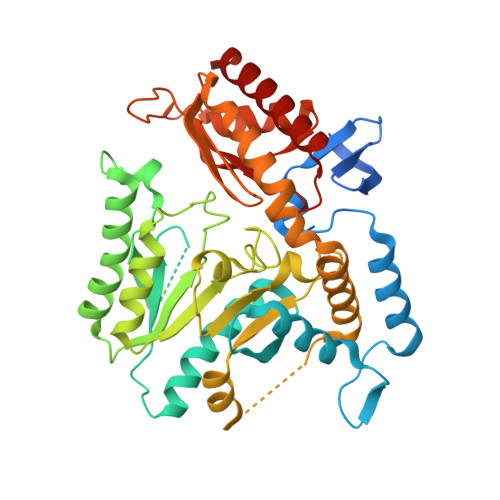
Characterization of a Putrescine Transaminase FromPseudomonas putidaand its Application to the Synthesis of Benzylamine Derivatives.
Galman, J.L., Gahloth, D., Parmeggiani, F., Slabu, I., Leys, D., Turner, N.J.(2018) Front Bioeng Biotechnol 6: 205-205
- PubMed: 30622946 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fbioe.2018.00205
- Primary Citation Related Structures:
6HX9 - PubMed Abstract:
The reductive amination of prochiral ketones using biocatalysts has been of great interest to the pharmaceutical industry in the last decade for integrating novel strategies in the production of chiral building blocks with the intent of minimizing impact on the environment. Amongst the enzymes able to catalyze the direct amination of prochiral ketones, pyridoxal 5'-phosphate (PLP) dependent ω-transaminases have shown great promise as versatile industrial biocatalysts with high selectivity, regioselectivity, and broad substrate scope. Herein the biochemical characterization of a putrescine transaminase from Pseudomonas putida (Pp-SpuC) was performed, which showed an optimum pH and temperature of 8.0 and 60°C, respectively. To gain further structural insight of this enzyme, we crystallized the protein in the apo form and determined the structure to 2.1 Å resolution which revealed a dimer that adopts a class I transaminase fold comparable to other class III transaminases. Furthermore we exploited its dual substrate recognition for biogenic diamines (i.e., cadaverine) and readily available monoamines (i.e., isopropylamine) for the synthesis of benzylamine derivatives with excellent product conversions and extremely broad substrate tolerance.
- School of Chemistry, Manchester Institute of Biotechnology, The University of Manchester, Manchester, United Kingdom.
Organizational Affiliation: