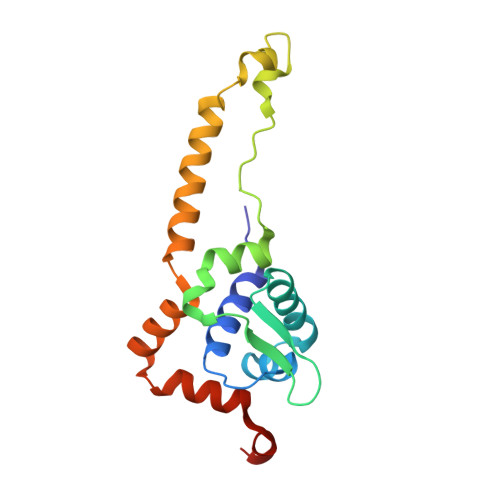
Crystal Structure of the Transcription Regulator RsrR Reveals a [2Fe-2S] Cluster Coordinated by Cys, Glu, and His Residues.
Volbeda, A., Martinez, M.T.P., Crack, J.C., Amara, P., Gigarel, O., Munnoch, J.T., Hutchings, M.I., Darnault, C., Le Brun, N.E., Fontecilla-Camps, J.C.(2019) J Am Chem Soc 141: 2367-2375
- PubMed: 30657661 Search on PubMed
- DOI: https://doi.org/10.1021/jacs.8b10823
- Primary Citation Related Structures:
6HSD, 6HSE, 6HSM - PubMed Abstract:
The recently discovered Rrf2 family transcriptional regulator RsrR coordinates a [2Fe-2S] cluster. Remarkably, binding of the protein to RsrR-regulated promoter DNA sequences is switched on and off through the facile cycling of the [2Fe-2S] cluster between +2 and +1 states. Here, we report high resolution crystal structures of the RsrR dimer, revealing that the [2Fe-2S] cluster is asymmetrically coordinated across the RsrR monomer-monomer interface by two Cys residues from one subunit and His and Glu residues from the other. To our knowledge, this is the first example of a protein bound [Fe-S] cluster with three different amino acid side chains as ligands, and of Glu acting as ligand to a [2Fe-2S] cluster. Analyses of RsrR structures revealed a conformational change, centered on Trp9, which results in a significant shift in the DNA-binding helix-turn-helix region.
- Université Grenoble Alpes, CEA, CNRS , IBS , Metalloproteins Unit, F-38044 Grenoble , France.
Organizational Affiliation: