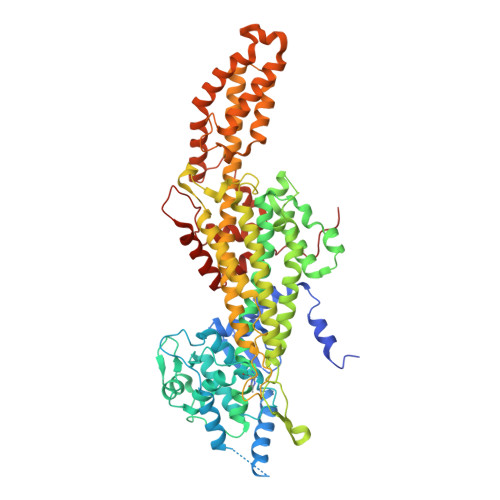
Substrate Tunnel Engineering Aided by X-ray Crystallography and Functional Dynamics Swaps the Function of MIO-Enzymes
Bata, Z., Molnar, Z., Madaras, E., Molnar, B., Santa-Bell, E., Varga, A., Leveles, I., Qian, R., Hammerschmidt, F., Paizs, C., Vertessy, B.G., Poppe, L.(2021) ACS Catal : 4538-4549