AURKC INCENP complex bound to BRD-7880
Abdul Azeez, K.R., Elkins, J.M.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Aurora kinase C | 274 | Homo sapiens | Mutation(s): 0 Gene Names: AURKC, AIE2, AIK3, AIRK3, ARK3, STK13 EC: 2.7.11.1 | 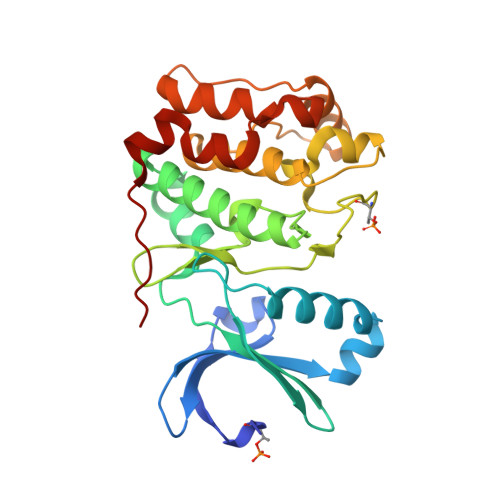 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9UQB9 GTEx: ENSG00000105146 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9UQB9 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Inner centromere protein | 70 | Homo sapiens | Mutation(s): 0 Gene Names: INCENP | 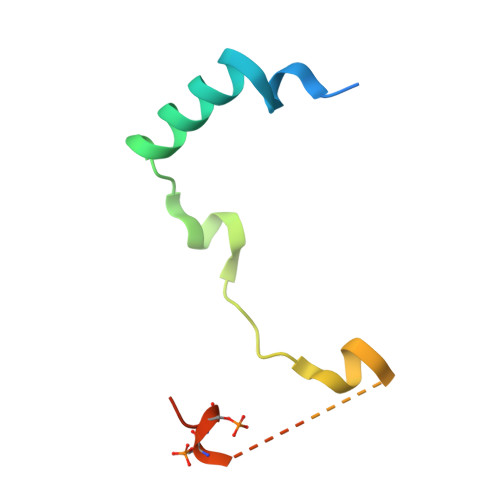 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q9NQS7 GTEx: ENSG00000149503 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9NQS7 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| VX6 Download:Ideal Coordinates CCD File | C [auth A] | CYCLOPROPANECARBOXYLIC ACID {4-[4-(4-METHYL-PIPERAZIN-1-YL)-6-(5-METHYL-2H-PYRAZOL-3-YLAMINO)-PYRIMIDIN-2-YLSULFANYL]-PHENYL}-AMIDE C23 H28 N8 O S GCIKSSRWRFVXBI-UHFFFAOYSA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | D [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| TPO Query on TPO | A | L-PEPTIDE LINKING | C4 H10 N O6 P |  | THR |
| SEP Query on SEP | B | L-PEPTIDE LINKING | C3 H8 N O6 P |  | SER |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 82.617 | α = 90 |
| b = 82.617 | β = 90 |
| c = 121 | γ = 120 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |