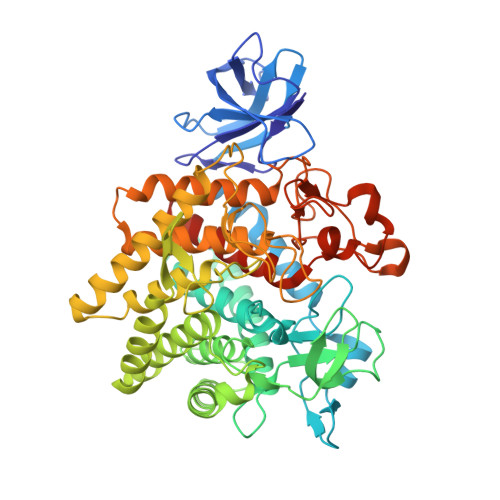
Structure of the GH9 glucosidase/glucosaminidase from Vibrio cholerae.
Wu, L., Davies, G.J.(2018) Acta Crystallogr F Struct Biol Commun 74: 512-523
- PubMed: 30084401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18011019
- Primary Citation Related Structures:
6GDT - PubMed Abstract:
Glycoside hydrolase family 9 (GH9) of carbohydrate-processing enzymes primarily consists of inverting endoglucanases. A subgroup of GH9 enzymes are believed to act as exo-glucosidases or exo-glucosaminidases, with many being found in organisms of the family Vibrionaceae, where they are proposed to function within the chitin-catabolism pathway. Here, it is shown that the GH9 enzyme from the pathogen Vibrio cholerae (hereafter referred to as VC0615) is active on both chitosan-derived and β-glucoside substrates. The structure of VC0615 at 3.17 Å resolution is reported from a crystal form with poor diffraction and lattice disorder. VC0615 was highly refractory to crystallization efforts, with crystals only appearing using a high protein concentration under conditions containing the precipitant poly-γ-glutamic acid (PGA). The structure is highly mobile within the crystal lattice, which is likely to reflect steric clashes between symmetry molecules which destabilize crystal packing. The overall tertiary structure of VC0615 is well resolved even at 3.17 Å resolution, which has allowed the structural basis for the exo-glucosidase/glucosaminidase activity of this enzyme to be investigated.
- York Structural Biology Laboratory, Department of Chemistry, The University of York, York YO10 5DD, England.
Organizational Affiliation: