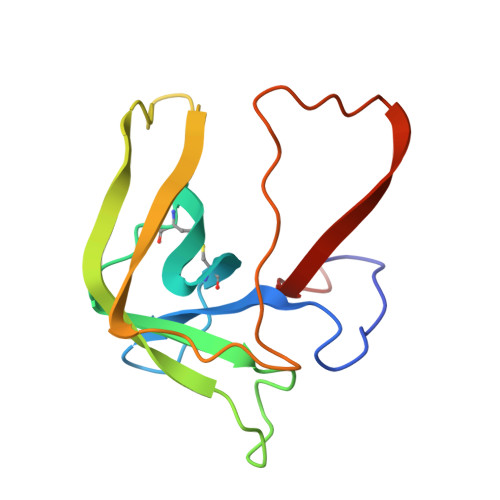
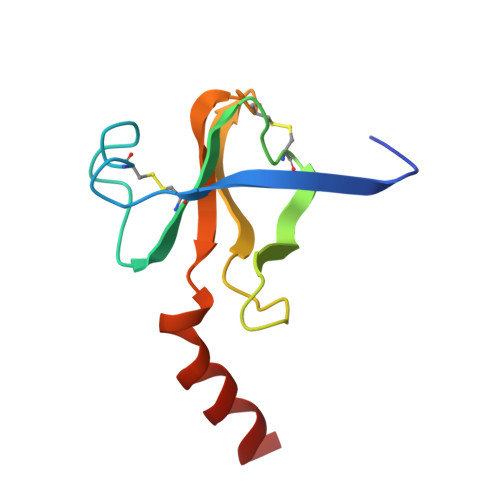
Structure of chymotrypsin-trifluoromethyl ketone inhibitor complexes: comparison of slowly and rapidly equilibrating inhibitors.
Brady, K., Wei, A.Z., Ringe, D., Abeles, R.H.(1990) Biochemistry 29: 7600-7607
- PubMed: 2271520 Search on PubMed
- DOI: https://doi.org/10.1021/bi00485a009
- Primary Citation Related Structures:
6GCH, 7GCH - PubMed Abstract:
The peptidyl trifluoromethyl ketones Ac-Phe-CF3 (1) and Ac-Leu-Phe-CF3 (2) are inhibitors of chymotrypsin. They differ in Ki (20 and 2 microM, respectively) as well as in their kinetics of association with chymotrypsin in that 1 is rapidly equilibrating, with an association rate too fast to be observed by steady-state techniques, while 2 is "slow binding", as defined by Morrison and Walsh [Morrison, J. F., & Walsh, C. T. (1988) Adv. Enzymol. Relat. Areas Mol. Biol. 61, 202], with a second-order association rate constant of 750 M-1 s-1 at pH 7.0 [Imperiali, B., & Abeles, R. (1986) Biochemistry 25, 3760]. The crystallographic structures of the complexes of gamma-chymotrypsin with inhibitors 1 and 2 have been determined in order to establish whether structural or conformational differences can be found which account for different kinetic and thermodynamic properties of the two inhibitors. In both complexes, the active-site Ser 195 hydroxyl forms a covalent hemiketal adduct with the trifluoromethyl ketone moiety of the inhibitor. In both complexes, the trifluoromethyl group is partially immobilized, but differences are observed in the degree of interaction of fluorine atoms with the active-site His 57 imidazole ring, with amide nitrogen NH 193, and with other portions of the inhibitor molecule. The enhanced potency of Ac-Leu-Phe-CF3 relative to Ac-Phe-CF3 is accounted for by van der Waals interactions of the leucine side chain of the inhibitor with His 57 and Ile 99 side chains and by a hydrogen bond of the acetyl terminus with amide NH 216 of the enzyme.(ABSTRACT TRUNCATED AT 250 WORDS)
- Department of Toxicology, Harvard School of Public Health, Boston, Massachusetts 02115.
Organizational Affiliation: