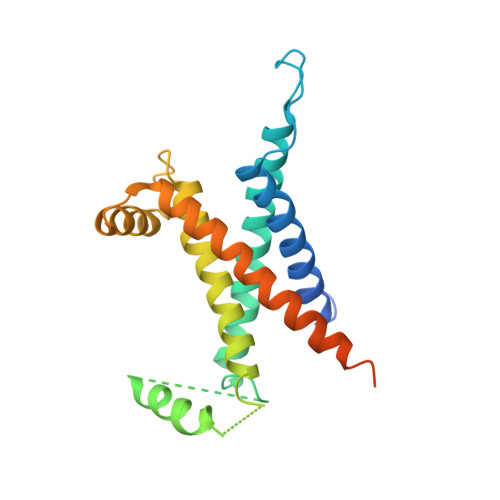
X-ray structural, functional and computational studies of the O2-sensitive E. coli hydrogenase-1 C19G variant reveal an unusual [4Fe-4S] cluster.
Volbeda, A., Mouesca, J.M., Darnault, C., Roessler, M.M., Parkin, A., Armstrong, F.A., Fontecilla-Camps, J.C.(2018) Chem Commun (Camb) 54: 7175-7178
- PubMed: 29888350 Search on PubMed
- DOI: https://doi.org/10.1039/c8cc02896f
- Primary Citation Related Structures:
6G94 - PubMed Abstract:
The crystal structure of the Escherichia coli O2-sensitive C19G [NiFe]-hydrogenase-1 variant shows that the mutation results in a novel FeS cluster, proximal to the Ni-Fe active site. While the proximal cluster of the native O2-tolerant enzyme can transfer two electrons to that site, EPR spectroscopy shows that the modified cluster can transfer only one electron, this shortfall coinciding with O2 sensitivity. Computational studies on electron transfer help to explain how the structural and redox properties of the novel FeS cluster modulate the observed phenotype.
- Univ. Grenoble Alpes, CEA, CNRS, IBS, F-38000 Grenoble, France. anne.volbeda@ibs.fr.
Organizational Affiliation: