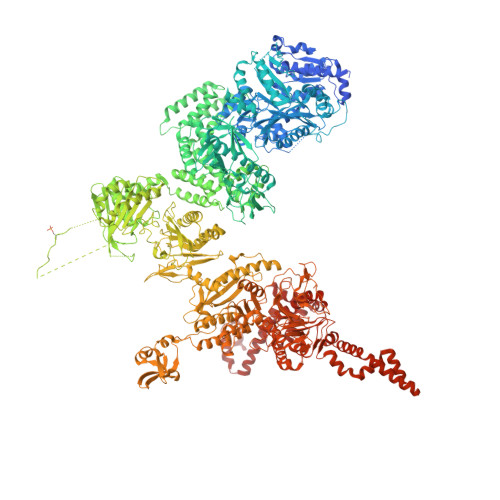
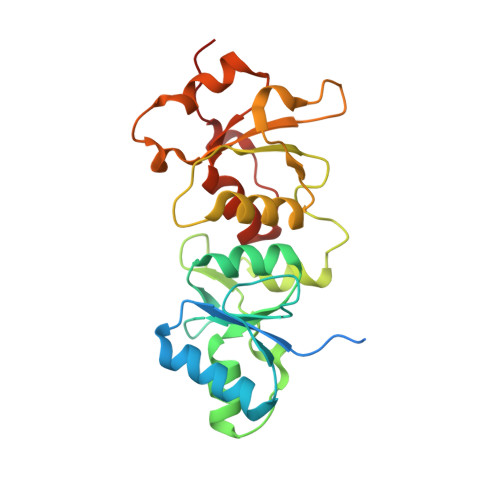
Structural basis for regulation of human acetyl-CoA carboxylase.
Hunkeler, M., Hagmann, A., Stuttfeld, E., Chami, M., Guri, Y., Stahlberg, H., Maier, T.(2018) Nature 558: 470-474
- PubMed: 29899443 Search on PubMed
- DOI: https://doi.org/10.1038/s41586-018-0201-4
- Primary Citation Related Structures:
6G2D, 6G2H, 6G2I - PubMed Abstract:
Acetyl-CoA carboxylase catalyses the ATP-dependent carboxylation of acetyl-CoA, a rate-limiting step in fatty acid biosynthesis 1,2 . Eukaryotic acetyl-CoA carboxylases are large, homodimeric multienzymes. Human acetyl-CoA carboxylase occurs in two isoforms: the metabolic, cytosolic ACC1, and ACC2, which is anchored to the outer mitochondrial membrane and controls fatty acid β-oxidation 1,3 . ACC1 is regulated by a complex interplay of phosphorylation, binding of allosteric regulators and protein-protein interactions, which is further linked to filament formation 1,4-8 . These filaments were discovered in vitro and in vivo 50 years ago 7,9,10 , but the structural basis of ACC1 polymerization and regulation remains unknown. Here, we identify distinct activated and inhibited ACC1 filament forms. We obtained cryo-electron microscopy structures of an activated filament that is allosterically induced by citrate (ACC-citrate), and an inactivated filament form that results from binding of the BRCT domains of the breast cancer type 1 susceptibility protein (BRCA1). While non-polymeric ACC1 is highly dynamic, filament formation locks ACC1 into different catalytically competent or incompetent conformational states. This unique mechanism of enzyme regulation via large-scale conformational changes observed in ACC1 has potential uses in engineering of switchable biosynthetic systems. Dissecting the regulation of acetyl-CoA carboxylase opens new paths towards counteracting upregulation of fatty acid biosynthesis in disease.
- Biozentrum, University of Basel, Basel, Switzerland. moritz_hunkeler@dfci.harvard.edu.
Organizational Affiliation: