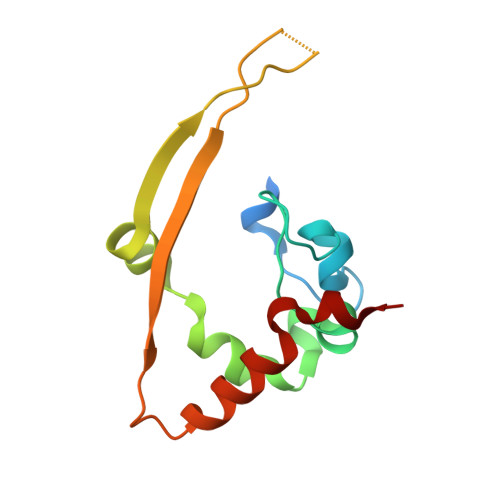
Structure and oligomerization state of the C-terminal region of the Middle East respiratory syndrome coronavirus nucleoprotein.
Nguyen, T.H.V., Lichiere, J., Canard, B., Papageorgiou, N., Attoumani, S., Ferron, F., Coutard, B.(2019) Acta Crystallogr D Struct Biol 75: 8-15
- PubMed: 30644840 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798318014948
- Primary Citation Related Structures:
6G13 - PubMed Abstract:
Middle East respiratory syndrome coronavirus (MERS-CoV) is a human pathogen responsible for a severe respiratory illness that emerged in 2012. Structural information about the proteins that constitute the viral particle is scarce. In order to contribute to a better understanding of the nucleoprotein (N) in charge of RNA genome encapsidation, the structure of the C-terminal domain of N from MERS-CoV obtained using single-crystal X-ray diffraction is reported here at 1.97 Å resolution. The molecule is present as a dimer in the crystal structure and this oligomerization state is confirmed in solution, as measured by additional methods including small-angle X-ray scattering measurements. Comparisons with the structures of the C-terminal domains of N from other coronaviruses reveals a high degree of structural conservation despite low sequence conservation, and differences in electrostatic potential at the surface of the protein.
- Aix-Marseille Université, AFMB UMR 7257, 13288 Marseilles, France.
Organizational Affiliation: