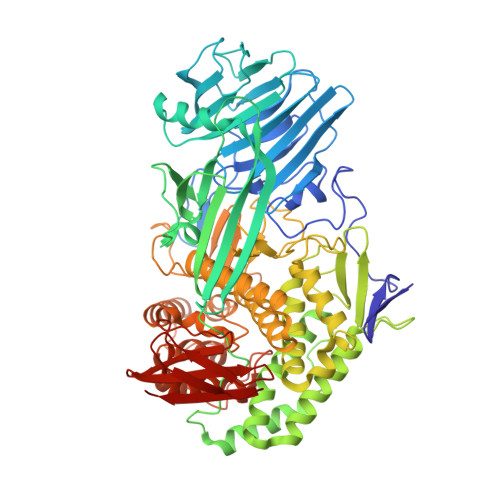
Novel insights into the degradation of beta-1,3-glucans by the cellulosome of Clostridium thermocellum revealed by structure and function studies of a family 81 glycoside hydrolase.
Kumar, K., Correia, M.A.S., Pires, V.M.R., Dhillon, A., Sharma, K., Rajulapati, V., Fontes, C.M.G.A., Carvalho, A.L., Goyal, A.(2018) Int J Biol Macromol 117: 890-901
- PubMed: 29870811 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2018.06.003
- Primary Citation Related Structures:
6FOP - PubMed Abstract:
The family 81 glycoside hydrolase (GH81) from Clostridium thermocellum is a β-1,3-glucanase belonging to cellulosomal complex. The gene encoding GH81 from Clostridium thermocellum (CtLam81A) was cloned and expressed displaying a molecular mass of ~82 kDa. CtLam81A showed maximum activity against laminarin (100 U/mg), followed by curdlan (65 U/mg), at pH 7.0 and 75 °C. CtLam81A displayed K m , 2.1 ± 0.12 mg/ml and V max , 109 ± 1.8 U/mg, against laminarin under optimized conditions. CtLam81A activity was significantly enhanced by Ca 2+ or Mg 2+ ions. Melting curve analysis of CtLam81A showed an increase in melting temperature from 91 °C to 96 °C by Ca 2+ or Mg 2+ ions and decreased to 82 °C by EDTA, indicating that Ca 2+ and Mg 2+ ions may be involved in catalysis and in maintaining structural integrity. TLC and MALDI-TOF analysis of β-1,3-glucan hydrolysed products released initially, showed β-1,3-glucan-oligosaccharides degree of polymerization (DP) from DP2 to DP7, confirming an endo-mode of action. The catalytically inactive mutant CtLam81A-E515A generated by site-directed mutagenesis was co-crystallized and tetragonal crystals diffracting up to 1.4 Å resolution were obtained. CtLam81A-E515A contained 15 α-helices and 38 β-strands forming a four-domain structure viz. a β-sandwich domain I at N-terminal, an α/β-domain II, an (α/α) 6 barrel domain III, and a small 5-stranded β-sandwich domain IV.
- Carbohydrate Enzyme Biotechnology Laboratory, Department of Bioscience and Bioengineering, Indian Institute of Technology Guwahati, Guwahati 781039, Assam, India.
Organizational Affiliation: