TbrPDEB1 structure with inhibitor NPD-1439
Salado, I.G., Moreno, C., Sakaine, G., Singh, A.K., Blaazer, A.R., Siderius, M., Matheeussen, A., Gul, S., Maes, L., Leurs, R., Brown, D.G., Augustyns, K.To be published.
Experimental Data Snapshot
Starting Model: other
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Phosphodiesterase | 360 | Trypanosoma brucei | Mutation(s): 0 Gene Names: PDEB1 EC: 3.1.4 | 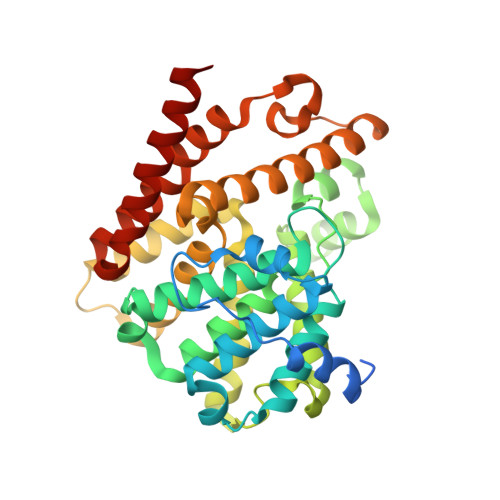 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q8WQX9 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| D68 Download:Ideal Coordinates CCD File | E [auth A] | 1-(2-{4-[(4aS,8aR)-4-[3,4-bis(difluoromethoxy)phenyl]-1-oxo-1,2,4a,5,8,8a-hexahydrophthalazin-2-yl]piperidin-1-yl}-2-oxoethyl)-4,4-dimethylpiperidine-2,6-dione C30 H34 F4 N4 O6 WVAQTVIALKSAMT-VQTJNVASSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | O [auth B], P [auth B], Q [auth B], V [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| ZN Download:Ideal Coordinates CCD File | F [auth A], R [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| GAI Download:Ideal Coordinates CCD File | C [auth A] D [auth A] L [auth A] M [auth A] N [auth A] | GUANIDINE C H5 N3 ZRALSGWEFCBTJO-UHFFFAOYSA-N |  | ||
| FMT Download:Ideal Coordinates CCD File | H [auth A], I [auth A], J [auth A], K [auth A] | FORMIC ACID C H2 O2 BDAGIHXWWSANSR-UHFFFAOYSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | G [auth A], S [auth B] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 115.53 | α = 90 |
| b = 114.82 | β = 108.25 |
| c = 68.31 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| REFMAC | phasing |